library(tidyverse)
library(mnormt)6 Monte Carlo Simulations
In this chapter, we use the same data-generating process described in Chapter 5 and run a series of Monte Carlo experiments. We consider two types of scenarios:
We change the distance between the means of the Gaussian distributions in groups 0 and 1. As this distance increases, the overlap between the two groups decreases.
We let the proportion of individuals in group 0 relative to group 1 vary. This simulates different levels of class imbalance.
Codes for graphical parameters
library(extrafont, quietly = TRUE)
col_group <- c("#00A08A","#F2AD00", "#1b95e0")
colour_methods <- c(
"OT" = "#CC79A7", "OT-M" = "#009E73",
"skh" = "darkgray",
"seq_1" = "#0072B2", "seq_2" = "#D55E00",
"fairadapt_1" = "#9966FF",
"fairadapt_2" = "#7CAE00"
)
colGpe1 <- col_group[2]
colGpe0 <- col_group[1]
colGpet <- col_group[3]
loadfonts(device = "pdf", quiet = TRUE)
font_size <- 20
font_family <- "CMU Serif"
path <- "./figs/"
if (!dir.exists(path)) dir.create(path)
source("../scripts/utils.R")\[ \definecolor{wongBlack}{RGB}{0,0,0} \definecolor{wongGold}{RGB}{230, 159, 0} \definecolor{wongLightBlue}{RGB}{86, 180, 233} \definecolor{wongGreen}{RGB}{0, 158, 115} \definecolor{wongYellow}{RGB}{240, 228, 66} \definecolor{wongBlue}{RGB}{0, 114, 178} \definecolor{wongOrange}{RGB}{213, 94, 0} \definecolor{wongPurple}{RGB}{204, 121, 167} \definecolor{colGpe1}{RGB}{0, 160, 138} \definecolor{colGpe0}{RGB}{242, 173, 0} \]
6.1 Functions
Let us redefine the function shown in Chapter 5 that will be used to perform the simulations here.
Functions used to generate the data (gen_data()), create counterfactuals with optimal transport (compute_ot_map(), apply_ot_transport()) and sequential transport (sequential_transport_12(), sequential_transport_21()), compute the total causal effect based on (causal_effects_cf())
## Data----
#' @param n0 Number of units in group 0.
#' @param n1 Number of units in group 1.
#' @param mu0 Mean of the two covariates in group 0.
#' @param mu1 Mean of the two covariates in group 1.
#' @param r0 Covariance of the two covariates in group 0.
#' @param r1 Covariance of the two covariates in group 1.
#' @param a Shift parameter for the mean in both groups (default to 1: no shift). Larger values decrease overlap.
#' @param seed Random seed for reproducibility.
gen_data <- function(n0 = 250,
n1 = 250,
mu0 = -1,
mu1 = +1,
r0 = +.7,
r1 = -.5,
a = 1,
seed = NULL) {
if (!is.null(seed)) set.seed(seed)
a0 <- 3
a1 <- 2
a2 <- -1.5
Mu0 <- rep(mu0, 2)
Mu1 <- rep(mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Generate covariates for each group
X0 <- rmnorm(n0, mean = a * Mu0, varcov = Sig0)
X1 <- rmnorm(n1, mean = a * Mu1, varcov = Sig1)
# Combine into a single covariate matrix
X <- rbind(X0, X1)
# Treatment indicator: 0 for first n0, 1 for next n1
A <- c(rep(0, n0), rep(1, n1))
# Random noise for each unit
E <- rnorm(n0 + n1)
# Outcomes
Y0 <- a1 * X[, 1] + a2 * X[, 2] + E
Y1 <- a1 * X[, 1] + a2 * X[, 2] + a0 + E
Y <- A * Y1 + (1 - A) * Y0
df <- tibble(
X1 = X[, 1],
X2 = X[, 2],
A = A,
Y0 = Y0,
Y1 = Y1,
Y = Y
)
df
}
## Optimal Transport----
#' Optimal transport mapping between two Gaussian distributions
#' (from \eqn{\mathcal{N}(\mu_{\text{source}}, \Sigma_{\text{source}})} to
#' \eqn{\mathcal{N}(\mu_{\text{target}}, \Sigma_{\text{target}})})
#'
#' @param mu_source Mean vector of the source Gaussian.
#' @param sigma_source Covariance matrix of the source Gaussian.
#' @param mu_target Mean vector of the target Gaussian.
#' @param sigma_target Covariance matrix of the target Gaussian.
compute_ot_map <- function(mu_source, sigma_source, mu_target, sigma_target) {
sqrt_sigma_source <- sqrtm(sigma_source)
sqrt_sigma_source_inv <- solve(sqrt_sigma_source)
inner <- sqrt_sigma_source %*% sigma_target %*% sqrt_sigma_source
sqrt_inner <- sqrtm(inner)
A <- sqrt_sigma_source_inv %*% sqrt_inner %*% sqrt_sigma_source_inv
list(A = A, shift = mu_target - A %*% mu_source)
}
#' Function to apply the transport map to simulated data
#'
#' @param X Observations to transport.
#' @param mapping Optimal transport mapping (from `compute_ot_map()`)?
apply_ot_transport <- function(X, mapping) {
A <- mapping$A
shift <- mapping$shift
t(apply(X, 1, function(x) as.vector(shift + A %*% x)))
}
## Penalized Transport----
#' @param X_source Matrix of observations to transport from the source group.
#' @param X_target Matrix of observations from the target group.
#' @param gamma A regularization parameter (default to 0.1).
transport_regul <- function(X_source,
X_target,
gamma) {
X_source <- as.matrix(X_source)
X_target <- as.matrix(X_target)
n_source <- nrow(X_source)
n_target <- nrow(X_target)
# Uniform weights
w_source <- rep(1 / n_source, n_source)
w_target <- rep(1 / n_target, n_target)
# Pairwise squared Euclidean distance
cost_mat <- as.matrix(dist(rbind(X_source, X_target)))^2
C <- cost_mat[1:n_source, (n_source + 1):(n_source + n_target)]
# Run Sinkhorn with entropic regularization gamma
skh_res <- T4transport::sinkhornD(
D = C, p = 2, wx = w_source, wy = w_target, lambda = gamma
)
# Extract and normalize plan
ot_plan_skh <- skh_res$plan
ot_plan_skh <- sweep(ot_plan_skh, 1, rowSums(ot_plan_skh), FUN = "/")
ot_plan_skh %*% X_target
}
## Transport Many-to-one----
#' @param X_source Source characteristics
#' @param X_target Target characteristics
#' @param method Algorithm to use for transport
transport_many_to_one <- function(X_source,
X_target,
method = "shortsimplex") {
n_source <- nrow(X_source)
n_target <- nrow(X_target)
# Uniform weights
w_source <- rep(1 / n_source, n_source)
w_target <- rep(1 / n_target, n_target)
# Cost matrix
cost <- as.matrix(dist(rbind(X_source, X_target)))
cost <- cost[1:n_source, (n_source + 1):(n_source + n_target)]
# Solve OT plan
ot_plan <- transport::transport(
w_source, w_target, costm = cost, method = method
)
# For each source unit, select the target with the highest mass
best_match <- ot_plan |>
dplyr::group_by(from) |>
dplyr::slice_max(mass, n = 1, with_ties = FALSE) |>
dplyr::ungroup()
# Matched matrix
X_matched <- X_target[best_match$to, , drop = FALSE]
X_matched
}
## Sequential Optimal Transport----
#' Sequential transport from N(M_source, S_source) to N(M_target, S_target),
#' along X1, then X2 | X1
#'
#' @param X n x 2 matrix of source observations.
#' @param M_source Mean vector of the source distribution (length 2).
#' @param S_source Covariance matrix of the source distribution (2x2).
#' @param M_target Mean vector of the target distribution.
#' @param S_target Covariance matrix of the target distribution.
sequential_transport_12 <- function(X,
M_source,
S_source,
M_target,
S_target) {
# marginal univariate transport along the first coordinate (X_1)
T1x <- qnorm(
pnorm(X[, 1], mean = M_source[1], sd = sqrt(S_source[1, 1])),
mean = M_target[1], sd = sqrt(S_target[1, 1])
)
# conditional parameters for X_2 | X_1
m_source <- M_source[2] + S_source[1, 2] / S_source[1, 1] * (X[, 1] - M_source[1])
s_source <- S_source[2, 2] - S_source[1, 2]^2 / S_source[1, 1]
m_target <- M_target[2] + S_target[1, 2] / S_target[1, 1] * (T1x - M_target[1])
s_target <- S_target[2, 2] - S_target[1, 2]^2 / S_target[1, 1]
# conditional transport for the second coordinate
T2x <- qnorm(
pnorm(X[, 2], mean = m_source, sd = sqrt(s_source)),
mean = m_target, sd = sqrt(s_target)
)
cbind(T1x, T2x)
}
#' Sequential transport from N(M_source, S_source) to N(M_target, S_target),
#' along X2, then X1 | X2
#'
#' @param X n x 2 matrix of source observations.
#' @param M_source Mean vector of the source distribution (length 2).
#' @param S_source Covariance matrix of the source distribution (2x2).
#' @param M_target Mean vector of the target distribution.
#' @param S_target Covariance matrix of the target distribution.
sequential_transport_21 <- function(X, M_source, S_source, M_target, S_target) {
# marginal univariate transport along X_2
T2x <- qnorm(
pnorm(X[, 2], mean = M_source[2], sd = sqrt(S_source[2, 2])),
mean = M_target[2], sd = sqrt(S_target[2, 2])
)
# conditional parameters for X_1 | X_2
m_source <- M_source[1] + S_source[1, 2] / S_source[2, 2] * (X[, 2] - M_source[2])
s_source <- S_source[1, 1] - S_source[1, 2]^2 / S_source[2, 2]
m_target <- M_target[1] + S_target[1, 2] / S_target[2, 2] * (T2x - M_target[2])
s_target <- S_target[1, 1] - S_target[1, 2]^2 / S_target[2, 2]
# conditional transport for X1 | X_2
T1x <- qnorm(
pnorm(X[, 1], mean = m_source, sd = sqrt(s_source)),
mean = m_target, sd = sqrt(s_target)
)
cbind(T1x, T2x)
}
fairadapt_12 <- function(df, quant_method = c("rangerQuants", "linearQuants")) {
adj_12 <- matrix(
c(0, 1, 1,
0, 0, 1,
0, 0, 0),
ncol = 3,
dimnames = rep(list(c("A","X1","X2")), 2),
byrow = TRUE
)
ind_0 <- which(df$A == 0)
data <- df |> select(A, X1, X2)
# A=0 (factuals) --> A=1 (counterfactuals)
data$A <- factor(data$A, levels = c("1", "0"))
fpt_model_0_to_1 <- fairadapt(
X2 ~ .,
train.data = data,
prot.attr = "A", adj.mat = adj_12,
quant.method = ifelse(quant_method == "rangerQuants", rangerQuants, linearQuants)
)
adapt_df_0 <- adaptedData(fpt_model_0_to_1)
# A=0 (factuals) --> A=1 (counterfactuals)
data$A <- factor(data$A, levels = c("0", "1"))
fpt_model_1_to_0 <- fairadapt(
X2 ~ .,
train.data = data,
prot.attr = "A", adj.mat = adj_12,
quant.method = ifelse(quant_method == "rangerQuants", rangerQuants, linearQuants)
)
adapt_df_1 <- adaptedData(fpt_model_1_to_0)
list(
adapt_df_0 = adapt_df_0[ind_0, !(names(adapt_df_0) %in% "A")],
adapt_df_1 = adapt_df_1[-ind_0, !(names(adapt_df_1) %in% "A")]
)
}
fairadapt_21 <- function(df, quant_method = c("rangerQuants", "linearQuants")) {
adj_21 <- matrix(
c(0, 1, 1,
0, 0, 0,
0, 1, 0),
ncol = 3,
dimnames = rep(list(c("A","X1","X2")), 2),
byrow = TRUE
)
ind_0 <- which(df$A == 0)
data <- df |> select(A, X1, X2)
# A=0 (factuals) --> A=1 (counterfactuals)
data$A <- factor(data$A, levels = c("1", "0"))
fpt_model_0_to_1 <- fairadapt(
X1 ~ .,
train.data = data,
prot.attr = "A", adj.mat = adj_21,
quant.method = ifelse(quant_method == "rangerQuants", rangerQuants, linearQuants)
)
adapt_df_0 <- adaptedData(fpt_model_0_to_1)
# A=0 (factuals) --> A=1 (counterfactuals)
data$A <- factor(data$A, levels = c("0", "1"))
fpt_model_1_to_0 <- fairadapt(
X1 ~ .,
train.data = data,
prot.attr = "A", adj.mat = adj_21,
quant.method = ifelse(quant_method == "rangerQuants", rangerQuants, linearQuants)
)
adapt_df_1 <- adaptedData(fpt_model_1_to_0)
list(
adapt_df_0 = adapt_df_0[ind_0, !(names(adapt_df_0) %in% "A")],
adapt_df_1 = adapt_df_1[-ind_0, !(names(adapt_df_1) %in% "A")]
)
}
## Causal Effect----
#' Estimation of total causal effect using counterfactuals.
#'
#' @param data_untreated Dataset with the untreated units only.
#' @param data_treated Dataset with the treated units only.
#' @param data_cf_untreated Counterfactuals for untreated had they been treated.
#' @param data_cf_treated Counterfactuals for treated had they been untreated.
#' @param Y_name Name of the column with the outcome variable.
#' @param A_name Name of the column with the treatment variable.
#' @param A_untreated Value of the treatment for the untreated units.
#'
#' @returns A list:
#' - `delta_0_i`: \eqn{\delta_(0)}, individual causal mediation effects for
#' \eqn{a=0} (computed on untreated),
#' - `delta_0`: \eqn{\bar{\delta}(0)}, average causal mediation effect for
#' \eqn{a=0} (computed on untreated),
#' - `delta_1_i`: \eqn{\delta_(1)}, individual causal mediation effects for
#' \eqn{a=1} (computed on treated),
#' - `delta_1`: \eqn{\bar{\delta}(1)}, average causal mediation effect for
#' \eqn{a=1} (computed on treated),
#' - `zeta_0_i`: \eqn{\zeta_(0)}, individual causal mediation effects for
#' \eqn{a=0} (computed on treaded),
#' - `zeta_0`: \eqn{\bar{\zeta}(0)}, average causal mediation effect for
#' \eqn{a=0} (computed on treated),
#' - `zeta_1_i`: \eqn{\zeta_(1)}, individual causal mediation effects for
#' \eqn{a=1} (computed on untreaded),
#' - `zeta_1`: \eqn{\bar{\zeta}(1)}, average causal mediation effect for
#' \eqn{a=1} (computed on untreated),
#' - `tot_effect`: \eqb{\tau}: average total effect (\eqn{\bar{\delta}(0) +
#' \bar{\zeta}(1)}).
#'
#' @importFrom randomForest randomForest
#' @importFrom dplyr pull select
#' @importFrom stats predict
#' @md
causal_effects_cf <- function(data_untreated,
data_treated,
data_cf_untreated,
data_cf_treated,
Y_name,
A_name,
A_untreated) {
n_untreated <- nrow(data_untreated)
n_treated <- nrow(data_treated)
# Outcome model for untreated
mu_untreated_model <- randomForest(
x = data_untreated |> dplyr::select(-!!Y_name, -!!A_name),
y = pull(data_untreated, !!Y_name)
)
# Outcome model for treated
mu_treated_model <- randomForest(
x = data_treated |> dplyr::select(-!!Y_name, -!!A_name),
y = pull(data_treated, !!Y_name)
)
# Observed outcome
y_untreated_obs <- data_untreated |> pull(!!Y_name)
y_treated_obs <- data_treated |> pull(!!Y_name)
# Natural Indirect Effect, using predictions
delta_0_i <- predict(mu_untreated_model, newdata = data_cf_untreated) -
predict(mu_untreated_model)
delta_0 <- mean(delta_0_i)
delta_1_i <- predict(mu_treated_model) -
predict(mu_treated_model, newdata = data_cf_treated)
delta_1 <- mean(delta_1_i)
# Natural Indirect Effect, using observed variables
delta_0_i_obs <- predict(mu_untreated_model, newdata = data_cf_untreated) -
y_untreated_obs
delta_0_obs <- mean(delta_0_i_obs)
delta_1_i_obs <- y_treated_obs -
predict(mu_treated_model, newdata = data_cf_treated)
delta_1_obs <- mean(delta_1_i_obs)
# Natural Direct Effect (only predictions)
zeta_0_i <- predict(mu_treated_model, newdata = data_cf_treated) -
predict(mu_untreated_model, newdata = data_cf_treated)
zeta_0 <- mean(zeta_0_i)
zeta_1_i <- predict(mu_treated_model, newdata = data_cf_untreated) -
predict(mu_untreated_model, newdata = data_cf_untreated)
zeta_1 <- mean(zeta_1_i)
# Total Causal Effect for treated
tot_effect <- delta_0 + zeta_1
tot_effect_obs <- delta_0_obs + zeta_1
list(
delta_0_i = delta_0_i,
delta_1_i = delta_1_i,
zeta_0_i = zeta_0_i,
zeta_1_i = zeta_1_i,
delta_0_i_obs = delta_0_i_obs,
delta_1_i_obs = delta_1_i_obs,
delta_0 = delta_0,
delta_1 = delta_1,
zeta_0 = zeta_0,
zeta_1 = zeta_1,
delta_0_obs = delta_0_obs,
delta_1_obs = delta_1_obs,
tot_effect = tot_effect,
tot_effect_obs = tot_effect_obs
)
}We define the function sim_f() to run a single replication of the Monte-Carlo simulations. This functions proceeds in the following steps:
- Generate data depending on the provided parameters for the DGP,
- Build Counterfactuals (with optimal transport and with sequential optimal transport) of units from the untreated group (\(A=0\)), as well as units from the treated group (\(A=1\)),
- Compute the total causal effect.
sim_f <- function(n0 = 250,
n1 = 250,
mu0,
mu1,
r0,
r1,
a,
seed = NULL) {
if (!is.null(seed)) set.seed(seed)
# 1. Generate data
df <- gen_data(
n0 = n0,
n1 = n1,
mu0 = mu0,
mu1 = mu1,
r0 = r0,
r1 = r1,
a = a,
seed = seed
)
# 2. Building Counterfactuals
## With Optimal Transport
# Transporting map for source: group 1, target: group 0 (careful here)
Sigma0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sigma1 <- matrix(c(1, r1, r1, 1), 2, 2)
Mu0 <- rep(a * mu0, 2)
Mu1 <- rep(a * mu1, 2)
# Mapping from group 0 to group 1
ot_map_0_to_1 <- compute_ot_map(
mu_source = Mu0, sigma_source = Sigma0,
mu_target = Mu1, sigma_target = Sigma1
)
# Mapping from group 1 to group 0
ot_map_1_to_0 <- compute_ot_map(
mu_source = Mu1, sigma_source = Sigma0,
mu_target = Mu0, sigma_target = Sigma0
)
# Apply transport map to treated units (A = 1)
X0 <- as.matrix(df[df$A == 0, c("X1", "X2")])
X1 <- as.matrix(df[df$A == 1, c("X1", "X2")])
X0_t <- apply_ot_transport(X = X0, mapping = ot_map_0_to_1)
colnames(X0_t) <- c(c("X1", "X2"))
X1_t <- apply_ot_transport(X = X1, mapping = ot_map_1_to_0)
colnames(X1_t) <- c(c("X1", "X2"))
# With OT-Matching
X0_tmatch <- transport_many_to_one(X_source = X0, X_target = X1)
X1_tmatch <- transport_many_to_one(X_source = X1, X_target = X0)
## With Entropy regularized transport
# Transport from group 0 to group 1:
X0_skh <- transport_regul(
X_source = X0,
X_target = X1,
gamma = 0.1
)
# Transport from group 1 to group 0:
X1_skh <- transport_regul(
X_source = X1,
X_target = X0,
gamma = 0.1
)
## With Sequential Transport
# Transport from group 0 to group 1: X1 then X2 | X1
X0_st_12 <- sequential_transport_12(
X = X0, M_source = Mu0, S_source = Sigma0, M_target = Mu1, S_target = Sigma1
)
# Transport from group 1 to group 0: X1 then X2 | X1
X1_st_12 <- sequential_transport_12(
X = X1, M_source = Mu1, S_source = Sigma1, M_target = Mu0, S_target = Sigma0
)
# Transport from group 0 to group 1: X2 then X1 | X2
X0_st_21 <- sequential_transport_21(
X = X0, M_source = Mu0, S_source = Sigma0, M_target = Mu1, S_target = Sigma1
)
# Transport from group 1 to group 0: X2 then X1 | X2
X1_st_21 <- sequential_transport_21(
X = X1, M_source = Mu1, S_source = Sigma1, M_target = Mu0, S_target = Sigma0
)
## With fairadapt RF
X_fairadapt_12_rf <- fairadapt_12(df = df, quant_method = "rangerQuants")
X_fairadapt_21_rf <- fairadapt_21(df = df, quant_method = "rangerQuants")
# Transport from group 0 to group 1: X1 then X2 | X1
X0_fairadapt_12_rf <- X_fairadapt_12_rf[[1]]
# Transport from group 1 to group 0: X1 then X2 | X1
X1_fairadapt_12_rf <- X_fairadapt_12_rf[[2]]
# Transport from group 0 to group 1: X2 then X1 | X2
X0_fairadapt_21_rf <- X_fairadapt_21_rf[[1]]
# Transport from group 1 to group 0: X2 then X1 | X2
X1_fairadapt_21_rf <- X_fairadapt_21_rf[[2]]
## With fairadapt linear
X_fairadapt_12_lin <- fairadapt_12(df = df, quant_method = "linearQuants")
X_fairadapt_21_lin <- fairadapt_21(df = df, quant_method = "linearQuants")
# Transport from group 0 to group 1: X1 then X2 | X1
X0_fairadapt_12_lin <- X_fairadapt_12_lin[[1]]
# Transport from group 1 to group 0: X1 then X2 | X1
X1_fairadapt_12_lin <- X_fairadapt_12_lin[[2]]
# Transport from group 0 to group 1: X2 then X1 | X2
X0_fairadapt_21_lin <- X_fairadapt_21_lin[[1]]
# Transport from group 1 to group 0: X2 then X1 | X2
X1_fairadapt_21_lin <- X_fairadapt_21_lin[[2]]
# 3. Measuring Total Causal Effect
tb <- df[, c("Y", "A", "X1", "X2")]
A_name <- "A"
A_untreated <- 0
Y_name <- "Y"
# Causal Mediation Analysis
med_mod_12 <- mediation::multimed(
outcome = "Y",
med.main = "X1",
med.alt = "X2",
treat = "A",
data = df
)
med_mod_21 <- mediation::multimed(
outcome = "Y",
med.main = "X2",
med.alt = "X1",
treat = "A",
data = df
)
delta_0_med <- mean((med_mod_12$d0.lb + med_mod_12$d0.ub) / 2) +
mean((med_mod_21$d0.lb + med_mod_21$d0.ub) / 2)
delta_1_med <- mean((med_mod_12$d1.lb + med_mod_12$d1.ub) / 2) +
mean((med_mod_21$d1.lb + med_mod_21$d1.ub) / 2)
tot_effect_med <- med_mod_12$tau
zeta_0_med <- tot_effect_med - delta_1_med
zeta_1_med <- tot_effect_med-delta_0_med
# With OT counterfactuals
tb_untreated <- tb |> filter(!!sym(A_name) == !!A_untreated)
tb_treated <- tb |> filter(!!sym(A_name) != !!A_untreated)
causal_effects_ot <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_t),
data_cf_treated = as_tibble(X1_t),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
# With OT-Matching counterfactuals
causal_effects_tmatch <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_tmatch),
data_cf_treated = as_tibble(X1_tmatch),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
# With entropy regularized transport
causal_effects_skh <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_skh),
data_cf_treated = as_tibble(X1_skh),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
# With Sequential Transport counterfactuals
causal_effect_sot_12 <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_st_12) |> magrittr::set_colnames(c("X1", "X2")),
data_cf_treated = as_tibble(X1_st_12) |> magrittr::set_colnames(c("X1", "X2")),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
causal_effect_sot_21 <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_st_21) |> magrittr::set_colnames(c("X1", "X2")),
data_cf_treated = as_tibble(X1_st_21) |> magrittr::set_colnames(c("X1", "X2")),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
# With fairadapt counterfactuals RF
causal_effect_fairadapt_12_rf <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_fairadapt_12_rf),
data_cf_treated = as_tibble(X1_fairadapt_12_rf),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
causal_effect_fairadapt_21_rf <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_fairadapt_21_rf),
data_cf_treated = as_tibble(X1_fairadapt_21_rf),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
# With fairadapt counterfactuals linear
causal_effect_fairadapt_12_lin <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_fairadapt_12_lin),
data_cf_treated = as_tibble(X1_fairadapt_12_lin),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
causal_effect_fairadapt_21_lin <- causal_effects_cf(
data_untreated = tb_untreated,
data_treated = tb_treated,
data_cf_untreated = as_tibble(X0_fairadapt_21_lin),
data_cf_treated = as_tibble(X1_fairadapt_21_lin),
Y_name = Y_name,
A_name = A_name,
A_untreated = A_untreated
)
tibble(
# Mediation
delta_0_med = delta_0_med,
delta_1_med = delta_1_med,
zeta_0_med = zeta_0_med,
zeta_1_med = zeta_1_med,
tot_effect_med = tot_effect_med,
# OT
delta_0_ot = causal_effects_ot$delta_0,
delta_1_ot = causal_effects_ot$delta_1,
delta_0_ot_obs = causal_effects_ot$delta_0_obs,
delta_1_ot_obs = causal_effects_ot$delta_1_obs,
zeta_0_ot = causal_effects_ot$zeta_0,
zeta_1_ot = causal_effects_ot$zeta_1,
tot_effect_ot = causal_effects_ot$tot_effect,
tot_effect_ot_obs = causal_effects_ot$tot_effect_obs,
# OT-M
delta_0_tmatch = causal_effects_tmatch$delta_0,
delta_1_tmatch = causal_effects_tmatch$delta_1,
delta_0_tmatch_obs = causal_effects_tmatch$delta_0_obs,
delta_1_tmatch_obs = causal_effects_tmatch$delta_1_obs,
zeta_0_tmatch = causal_effects_tmatch$zeta_0,
zeta_1_tmatch = causal_effects_tmatch$zeta_1,
tot_effect_tmatch = causal_effects_tmatch$tot_effect,
tot_effect_tmatch_obs = causal_effects_tmatch$tot_effect_obs,
# SKH
delta_0_skh = causal_effects_skh$delta_0,
delta_1_skh = causal_effects_skh$delta_1,
delta_0_skh_obs = causal_effects_skh$delta_0_obs,
delta_1_skh_obs = causal_effects_skh$delta_1_obs,
zeta_0_skh = causal_effects_skh$zeta_0,
zeta_1_skh = causal_effects_skh$zeta_1,
tot_effect_skh = causal_effects_skh$tot_effect,
tot_effect_skh_obs = causal_effects_skh$tot_effect_obs,
# SOT 12
delta_0_sot_12 = causal_effect_sot_12$delta_0,
delta_1_sot_12 = causal_effect_sot_12$delta_1,
delta_0_sot_12_obs = causal_effect_sot_12$delta_0_obs,
delta_1_sot_12_obs = causal_effect_sot_12$delta_1_obs,
zeta_0_sot_12 = causal_effect_sot_12$zeta_0,
zeta_1_sot_12 = causal_effect_sot_12$zeta_1,
tot_effect_sot_12 = causal_effect_sot_12$tot_effect,
tot_effect_sot_12_obs = causal_effect_sot_12$tot_effect_obs,
# SOT 21
delta_0_sot_21 = causal_effect_sot_21$delta_0,
delta_1_sot_21 = causal_effect_sot_21$delta_1,
delta_0_sot_21_obs = causal_effect_sot_21$delta_0_obs,
delta_1_sot_21_obs = causal_effect_sot_21$delta_1_obs,
zeta_0_sot_21 = causal_effect_sot_21$zeta_0,
zeta_1_sot_21 = causal_effect_sot_21$zeta_1,
tot_effect_sot_21 = causal_effect_sot_21$tot_effect,
tot_effect_sot_21_obs = causal_effect_sot_21$tot_effect_obs,
# fairadapt 12 RF
delta_0_fairadapt_12_rf = causal_effect_fairadapt_12_rf$delta_0,
delta_1_fairadapt_12_rf = causal_effect_fairadapt_12_rf$delta_1,
delta_0_fairadapt_12_obs_rf = causal_effect_fairadapt_12_rf$delta_0_obs,
delta_1_fairadapt_12_obs_rf = causal_effect_fairadapt_12_rf$delta_1_obs,
zeta_0_fairadapt_12_rf = causal_effect_fairadapt_12_rf$zeta_0,
zeta_1_fairadapt_12_rf = causal_effect_fairadapt_12_rf$zeta_1,
tot_effect_fairadapt_12_rf = causal_effect_fairadapt_12_rf$tot_effect,
tot_effect_fairadapt_12_obs_rf = causal_effect_fairadapt_12_rf$tot_effect_obs,
# fairadapt 21 RF
delta_0_fairadapt_21_rf = causal_effect_fairadapt_21_rf$delta_0,
delta_1_fairadapt_21_rf = causal_effect_fairadapt_21_rf$delta_1,
delta_0_fairadapt_21_obs_rf = causal_effect_fairadapt_21_rf$delta_0_obs,
delta_1_fairadapt_21_obs_rf = causal_effect_fairadapt_21_rf$delta_1_obs,
zeta_0_fairadapt_21_rf = causal_effect_fairadapt_21_rf$zeta_0,
zeta_1_fairadapt_21_rf = causal_effect_fairadapt_21_rf$zeta_1,
tot_effect_fairadapt_21_rf = causal_effect_fairadapt_21_rf$tot_effect,
tot_effect_fairadapt_21_obs_rf = causal_effect_fairadapt_21_rf$tot_effect_obs,
# fairadapt 12 linear
delta_0_fairadapt_12_lin = causal_effect_fairadapt_12_lin$delta_0,
delta_1_fairadapt_12_lin = causal_effect_fairadapt_12_lin$delta_1,
delta_0_fairadapt_12_obs_lin = causal_effect_fairadapt_12_lin$delta_0_obs,
delta_1_fairadapt_12_obs_lin = causal_effect_fairadapt_12_lin$delta_1_obs,
zeta_0_fairadapt_12_lin = causal_effect_fairadapt_12_lin$zeta_0,
zeta_1_fairadapt_12_lin = causal_effect_fairadapt_12_lin$zeta_1,
tot_effect_fairadapt_12_lin = causal_effect_fairadapt_12_lin$tot_effect,
tot_effect_fairadapt_12_obs_lin = causal_effect_fairadapt_12_lin$tot_effect_obs,
# fairadapt 21 linear
delta_0_fairadapt_21_lin = causal_effect_fairadapt_21_lin$delta_0,
delta_1_fairadapt_21_lin = causal_effect_fairadapt_21_lin$delta_1,
delta_0_fairadapt_21_obs_lin = causal_effect_fairadapt_21_lin$delta_0_obs,
delta_1_fairadapt_21_obs_lin = causal_effect_fairadapt_21_lin$delta_1_obs,
zeta_0_fairadapt_21_lin = causal_effect_fairadapt_21_lin$zeta_0,
zeta_1_fairadapt_21_lin = causal_effect_fairadapt_21_lin$zeta_1,
tot_effect_fairadapt_21_lin = causal_effect_fairadapt_21_lin$tot_effect,
tot_effect_fairadapt_21_obs_lin = causal_effect_fairadapt_21_lin$tot_effect_obs,
n0 = n0,
n1 = n1,
seed = seed,
mu0 = mu0,
mu1 = mu1,
r0 = r0,
r1 = r1,
a = a
)
}6.2 Varying the Distance Between the Means
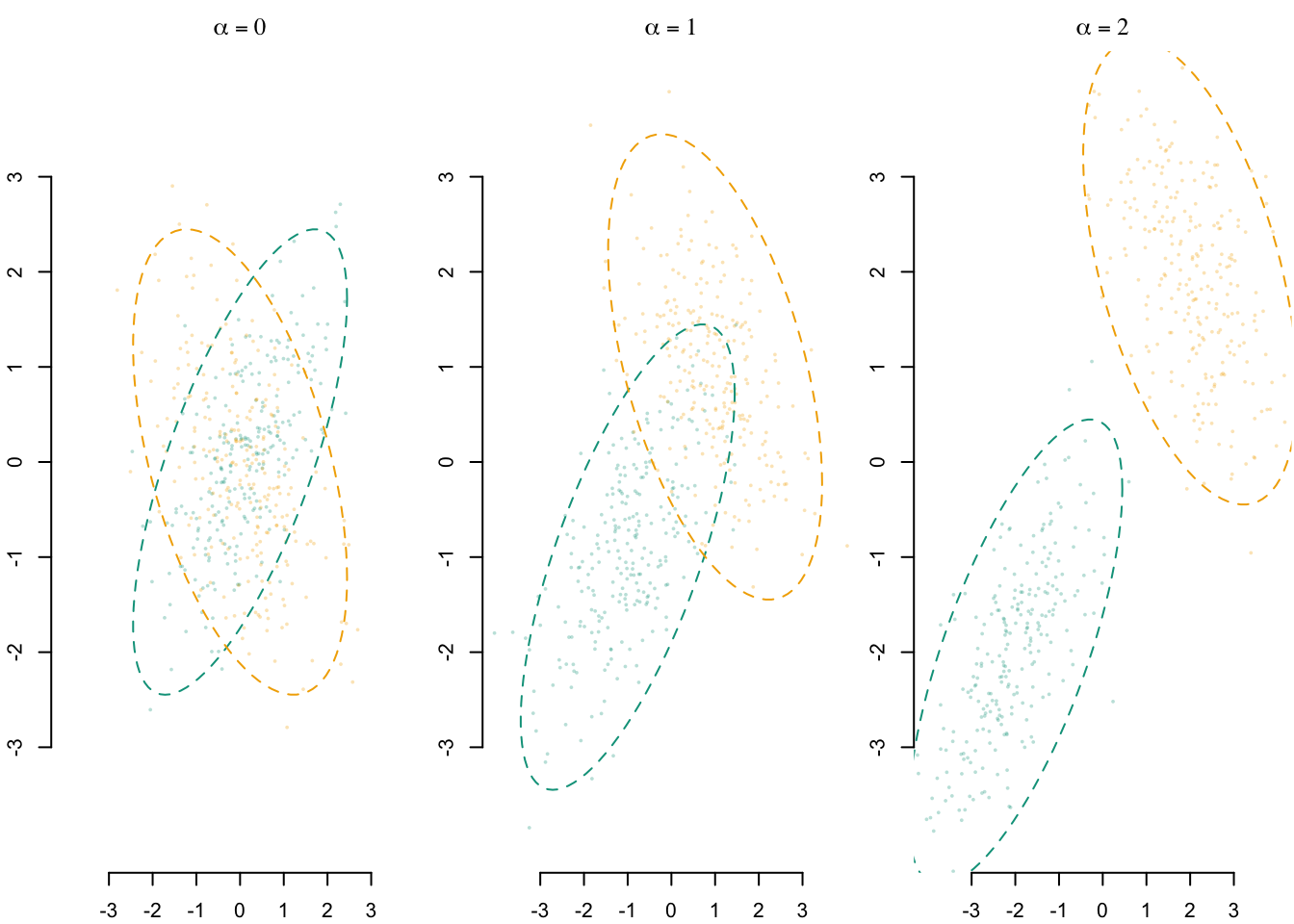
We make the distance between the means of the Gaussian distributions of the two groups increase. With \(\boldsymbol{\mu}_0 = \begin{pmatrix}-1\\-1\end{pmatrix}\) and \(\boldsymbol{\mu}_1 = \begin{pmatrix}1\\1\end{pmatrix}\), the distance is equal to \(1\). We apply a scalar coefficient \(\alpha\geq 0\) to the means to increase that distance: \(a\boldsymbol{\mu}_0\) and \(a\boldsymbol{\mu}_1\). We make \(\alpha\) vary from 0 to 2 by steps of .1. When \(\alpha=0\), the distance is equal to \(0\), when \(\alpha=2\), the distance is equal to \(4\).
A visual representation of the samples that can be drawn from the DGP with three values of \(\alpha\) is provided in Figure 6.1.
Codes to create the Figure.
library(tikzDevice)
export_tikz <- FALSE
scale <- 1.7
if (export_tikz == TRUE) {
filename <- "gaussian-example-alpha"
tikz(paste0("figs/", filename, ".tex"), width = scale*2.2, height = scale*1)
}
draw_ellipse <- function(mu,
sigma,
col = "black",
lty = 1,
lwd = 1,
level = 0.95,
...) {
angles <- seq(0, 2 * pi, length.out = 100)
vals <- sqrt(
qchisq(level, df = 2)) * t(chol(sigma)) %*% rbind(cos(angles), sin(angles)
)
lines(mu[1] + vals[1, ], mu[2] + vals[2, ], col = col, lty = lty, lwd = lwd, ...)
}
set.seed(123)
mu0 <- -1
mu1 <- +1
r0 <- +.7
r1 <- -.5
# Generate data with alpha=0, alpha=1, alpha=2
data_alpha_0 <- gen_data(a = 0, mu0 = mu0, mu1 = mu1, r0 = r0, r1 = r1)
data_alpha_1 <- gen_data(a = 1, mu0 = mu0, mu1 = mu1, r0 = r0, r1 = r1)
data_alpha_2 <- gen_data(a = 2, mu0 = mu0, mu1 = mu1, r0 = r0, r1 = r1)
par(mar = c(2.1, 2.1, 2.1, 0.1), mfrow = c(1, 3))
x_lim <- c(-4, 4)
y_lim <- c(-4, 4)
cex_pts <- .4
# With alpha=0----
a <- 0
x_lab <- "$\\alpha = 0$"
if (export_tikz == FALSE) x_lab <- latex2exp::TeX(x_lab)
plot(
data_alpha_0$X1[data_alpha_0$A == 0],
data_alpha_0$X2[data_alpha_0$A == 0],
pch = 16,
cex = cex_pts,
col = adjustcolor(colGpe0, alpha = .3),
xlim = x_lim, ylim = y_lim,
xlab = "", ylab = "",
main = x_lab,
family = font_family,
axes = FALSE
)
axis(1, at = -3:3)
axis(2, at = -3:3)
title(xlab = "X1", ylab="X2", line=2, cex.lab=1.2, family = font_family)
points(
data_alpha_0$X1[data_alpha_0$A == 1],
data_alpha_0$X2[data_alpha_0$A == 1],
col = adjustcolor(colGpe1, alpha = .3), pch = 16, cex = cex_pts
)
# True mean and covariance (scaled by 'a')
Mu0 <- rep(a * mu0, 2)
Mu1 <- rep(a * mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Add ellipses
draw_ellipse(Mu0, Sig0, col = colGpe0, lty = 2)
draw_ellipse(Mu1, Sig1, col = colGpe1, lty = 2)
# With alpha=1----
a <- 1
x_lab <- "$\\alpha = 1$"
if (export_tikz == FALSE) x_lab <- latex2exp::TeX(x_lab)
plot(
data_alpha_1$X1[data_alpha_1$A == 0],
data_alpha_1$X2[data_alpha_1$A == 0],
pch = 16, cex = cex_pts,
col = adjustcolor(colGpe0, alpha = .3),
xlim = x_lim, ylim = y_lim,
xlab = "", ylab = "",
main = x_lab,
family = font_family,
axes = FALSE
)
axis(1, at = -3:3)
axis(2, at = -3:3)
title(xlab = "X1", ylab="X2", line=2, cex.lab=1.2, family = font_family)
points(
data_alpha_1$X1[data_alpha_1$A == 1],
data_alpha_1$X2[data_alpha_1$A == 1],
col = adjustcolor(colGpe1, alpha = .3), pch = 16, cex = cex_pts
)
# True mean and covariance (scaled by 'a')
Mu0 <- rep(a * mu0, 2)
Mu1 <- rep(a * mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Add ellipses
draw_ellipse(Mu0, Sig0, col = colGpe0, lty = 2)
draw_ellipse(Mu1, Sig1, col = colGpe1, lty = 2)
# With alpha=2----
a <- 2
x_lab <- "$\\alpha = 2$"
if (export_tikz == FALSE) x_lab <- latex2exp::TeX(x_lab)
plot(
data_alpha_2$X1[data_alpha_2$A == 0],
data_alpha_2$X2[data_alpha_2$A == 0],
pch = 16, cex = cex_pts,
col = adjustcolor(colGpe0, alpha = .3),
xlim = x_lim, ylim = y_lim,
xlab = "", ylab = "",
main = x_lab,
family = font_family,
axes = FALSE
)
axis(1, at = -3:3)
axis(2, at = -3:3)
title(xlab = "X1", ylab="X2", line=2, cex.lab=1.2, family = font_family)
points(
data_alpha_2$X1[data_alpha_2$A == 1],
data_alpha_2$X2[data_alpha_2$A == 1],
col = adjustcolor(colGpe1, alpha = .3), pch = 16, cex = cex_pts
)
# True mean and covariance (scaled by 'a')
Mu0 <- rep(a * mu0, 2)
Mu1 <- rep(a * mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Add ellipses
draw_ellipse(Mu0, Sig0, col = colGpe0, lty = 2)
draw_ellipse(Mu1, Sig1, col = colGpe1, lty = 2)
if (export_tikz == TRUE) {
dev.off()
plot_to_pdf(
filename = filename, path = "./figs/", keep_tex = FALSE, crop = FALSE
)
}
We define a grid with the different simulations.
n_repl <- 200
grid_params <- expand_grid(
n0 = 250,
n1 = 250,
mu0 = -1,
mu1 = +1,
r0 = +.7,
r1 = -.5,
a = seq(0, 2, by = .1),
seed = seq_len(n_repl)
)The simulations can be run in parallel.
The codes to run the simulations
# This chunk takes about XX minutes to run.
# We do not evaluate when compiling the document.
# Instead, we load previously obtained results.
library(pbapply)
library(parallel)
ncl <- detectCores()-1
(cl <- makeCluster(ncl))
clusterEvalQ(cl, {
library(tidyverse)
library(mnormt)
library(expm)
library(randomForest)
library(fairadapt)
}) |>
invisible()
clusterExport(
cl = cl, c(
"gen_data", "compute_ot_map", "apply_ot_transport",
"transport_regul", "transport_many_to_one",
"sequential_transport_12", "sequential_transport_21",
"causal_effects_cf", "sim_f", "grid_params", "fairadapt_12",
"fairadapt_21"
)
)
# For an unknown reason, there seems to be an issue with performing this in
# one go. Let us divide the task into two.
res_sim_1 <- pbapply::pblapply(1:2100, function(i) {
grid_params_current <- grid_params[i, ]
n0 <- grid_params_current$n0
n1 <- grid_params_current$n1
mu0 <- grid_params_current$mu0
mu1 <- grid_params_current$mu1
r0 <- grid_params_current$r0
r1 <- grid_params_current$r1
a <- grid_params_current$a
seed <- grid_params_current$seed
sim_f(
n0 = n0, n1 = n1,
mu0 = mu0, mu1 = mu1,
r0 = r0, r1 = r1,
a = a, seed = seed
)
}, cl = cl)
res_sim_2 <- pbapply::pblapply(2101:nrow(grid_params), function(i) {
grid_params_current <- grid_params[i, ]
n0 <- grid_params_current$n0
n1 <- grid_params_current$n1
mu0 <- grid_params_current$mu0
mu1 <- grid_params_current$mu1
r0 <- grid_params_current$r0
r1 <- grid_params_current$r1
a <- grid_params_current$a
seed <- grid_params_current$seed
sim_f(
n0 = n0, n1 = n1,
mu0 = mu0, mu1 = mu1,
r0 = r0, r1 = r1,
a = a, seed = seed
)
}, cl = cl)
res_sim_1 <- list_rbind(res_sim_1)
res_sim_2 <- list_rbind(res_sim_2)
res_sim <- res_sim_1 |> bind_rows(res_sim_2)
stopCluster(cl)
save(res_sim, file = "../output/res_sim-gaussian-mc.rda")We load previously obtained results:
load("../output/res_sim-gaussian-mc.rda")Codes to create the Figure.
a1 <- 2
a2 <- -1.5
export_pdf <- FALSE
x_lab <- "$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"
y_lab <- "Distance to $\\bar\\tau$"
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
}
data_plot <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
tot_effect_med, tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
tot_effect_med, tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin
),
names_to = "type", values_to = "tau"
) |>
mutate(
type = factor(
type,
levels = c(
"tot_effect_med", "tot_effect_ot", "tot_effect_tmatch",
"tot_effect_skh", "tot_effect_sot_12", "tot_effect_sot_21",
"tot_effect_fairadapt_12_lin", "tot_effect_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_causal_effect = (a1+a2) * (a*mu1 - a*mu0) + 3 - tau
)
p <- ggplot(
data = data_plot,
mapping = aes(x = factor(a), y = dist_to_causal_effect)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none",
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
if (export_pdf == TRUE) {
filename <- "gaussian-mc-alpha-tau"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
width = 3.3, height = 3.5,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p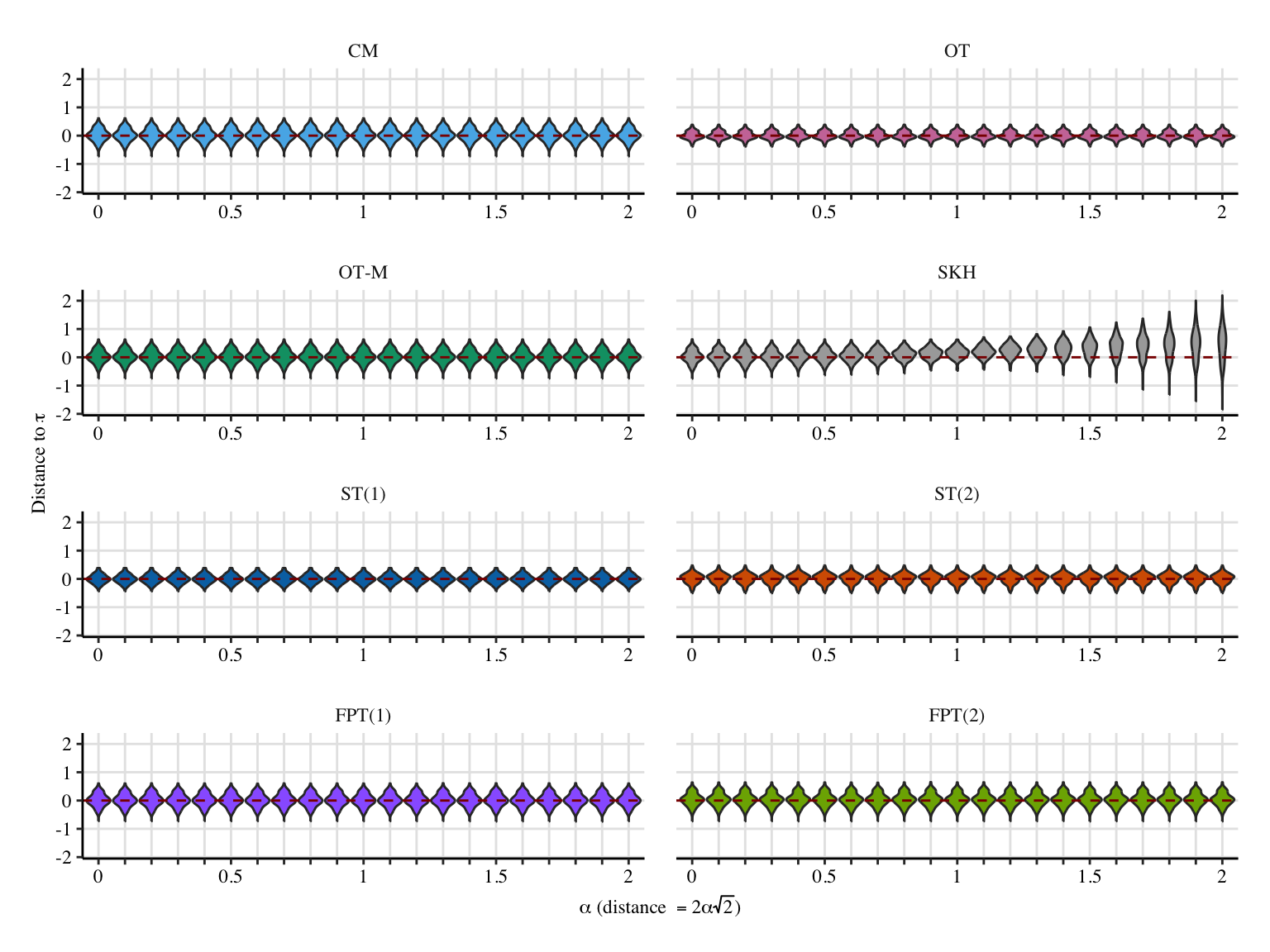
Codes to create the Figure.
export_pdf <- FALSE
x_lab <- "$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"
# y_lab <- "Distance to $\\bar\\delta(0)$"
y_lab <- "Distance to $\\bar\\delta$"
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
}
data_plot <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
delta_0_med, delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
delta_0_med, delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin
),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"delta_0_med", "delta_0_ot", "delta_0_tmatch",
"delta_0_skh", "delta_0_sot_12", "delta_0_sot_21",
"delta_0_fairadapt_12_lin", "delta_0_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = ((a1+a2)*(a*mu1-a*mu0)) - val
)
p <- ggplot(
data = data_plot,
mapping = aes(x = factor(a), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
if (export_pdf == TRUE) {
filename <- "gaussian-mc-alpha-delta0"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
width = 3.3, height = 3.5,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p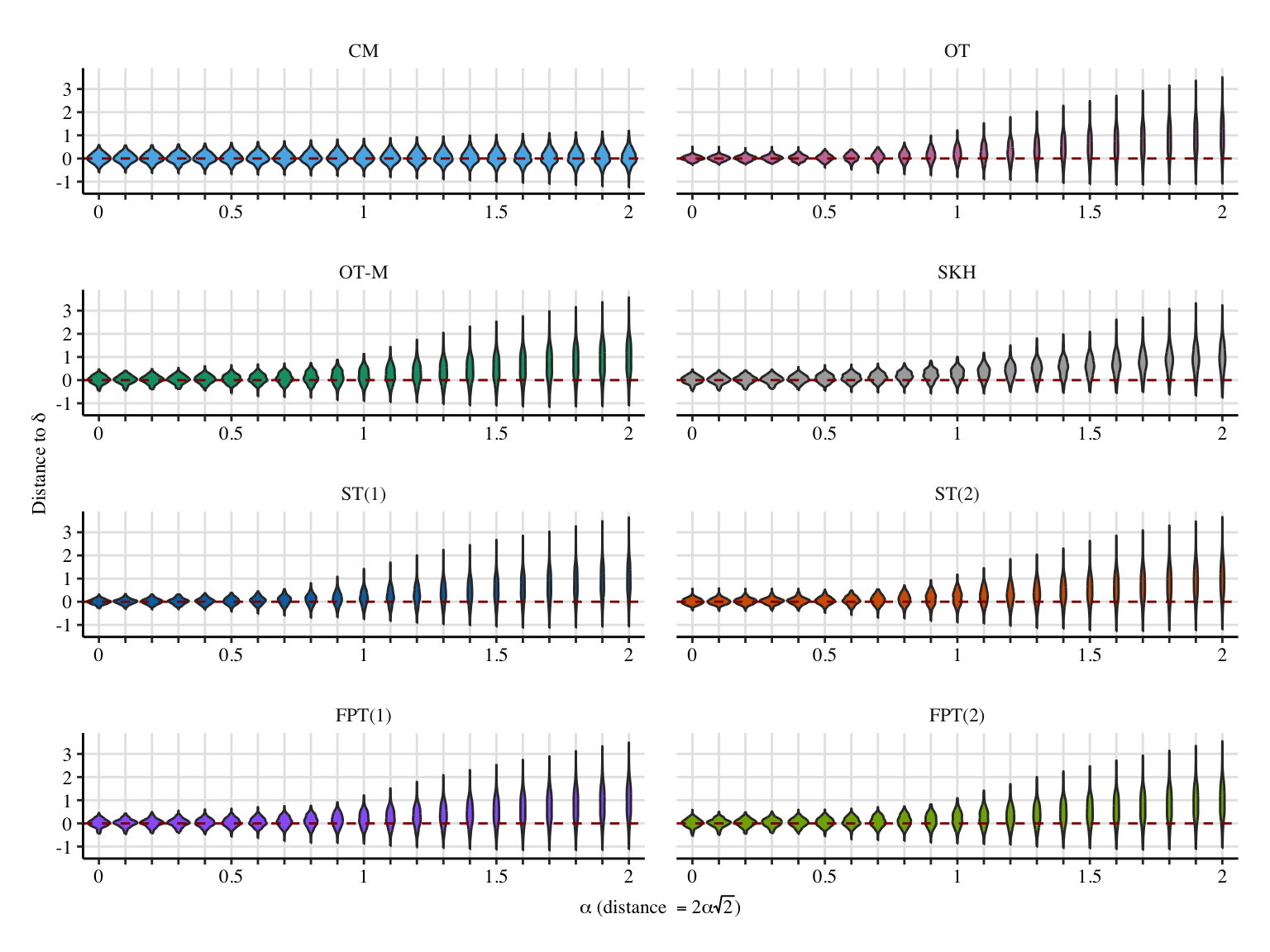
Codes to create the Figure.
data_plot <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
delta_1_med, delta_1_ot, delta_1_tmatch,
delta_1_skh, delta_1_sot_12, delta_1_sot_21,
delta_1_fairadapt_12_lin, delta_1_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
delta_1_med, delta_1_ot, delta_1_tmatch,
delta_1_skh, delta_1_sot_12, delta_1_sot_21,
delta_1_fairadapt_12_lin, delta_1_fairadapt_21_lin),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"delta_1_med", "delta_1_ot", "delta_1_tmatch",
"delta_1_skh", "delta_1_sot_12", "delta_1_sot_21",
"delta_1_fairadapt_12_lin", "delta_1_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = ((a1+a2)*(a*mu1-a*mu0)) - val
)
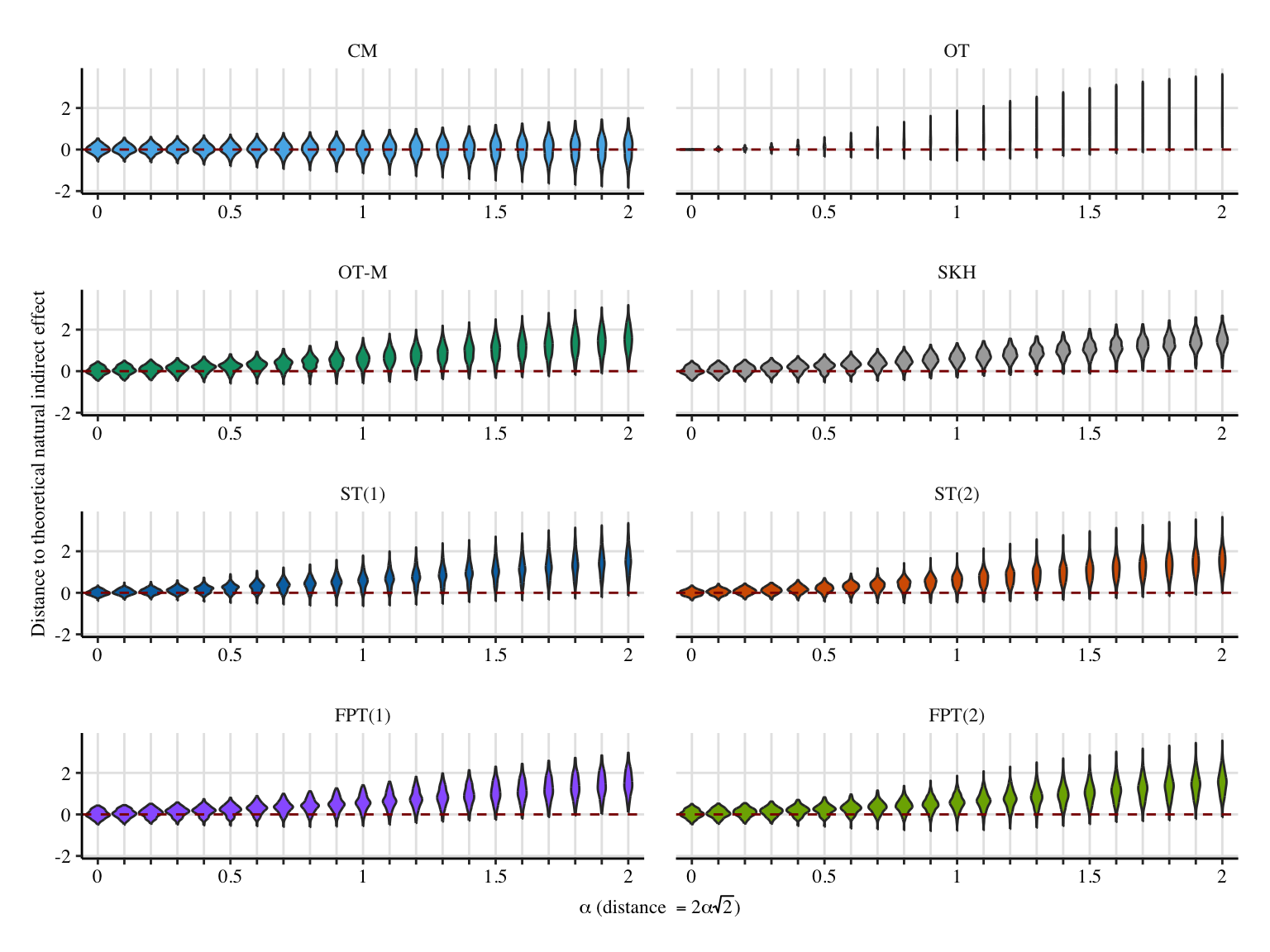
ggplot(
data = data_plot,
mapping = aes(x = factor(a), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
labs(
x = latex2exp::TeX("$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"),
y = "Distance to theoretical natural indirect effect",
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
Codes to create the Figure.
data_plot <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
zeta_0_med, zeta_0_ot, zeta_0_tmatch,
zeta_0_skh, zeta_0_sot_12, zeta_0_sot_21,
zeta_0_fairadapt_12_lin, zeta_0_fairadapt_21_lin
) |>
pivot_longer(
cols = c(zeta_0_med, zeta_0_ot, zeta_0_tmatch,
zeta_0_skh, zeta_0_sot_12, zeta_0_sot_21,
zeta_0_fairadapt_12_lin, zeta_0_fairadapt_21_lin),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"zeta_0_med", "zeta_0_ot", "zeta_0_tmatch",
"zeta_0_skh", "zeta_0_sot_12", "zeta_0_sot_21",
"zeta_0_fairadapt_12_lin", "zeta_0_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = 3 - val
)
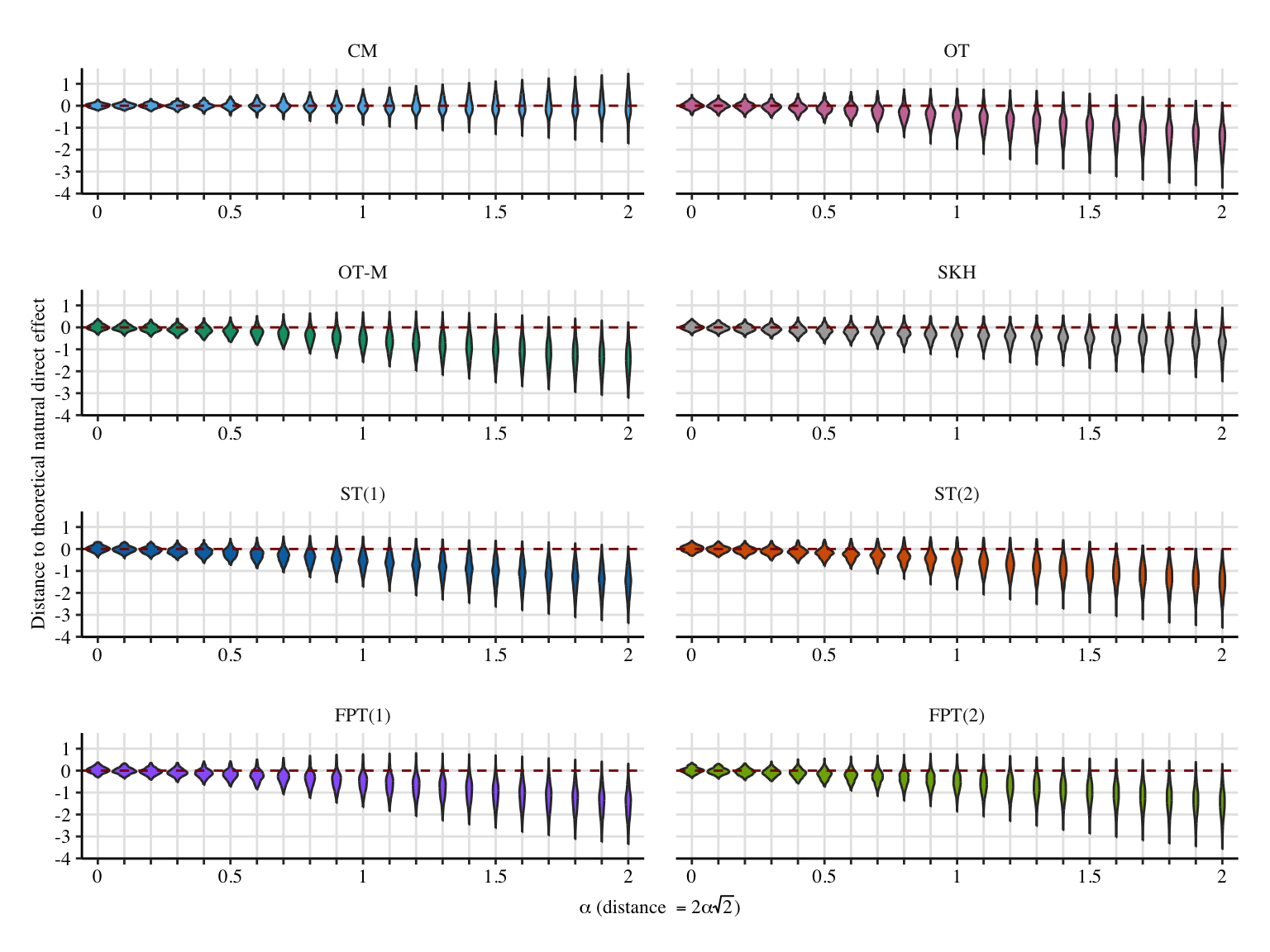
ggplot(
data = data_plot,
mapping = aes(x = factor(a), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
labs(
x = latex2exp::TeX("$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"),
y = "Distance to theoretical natural direct effect",
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
Codes to create the Figure.
p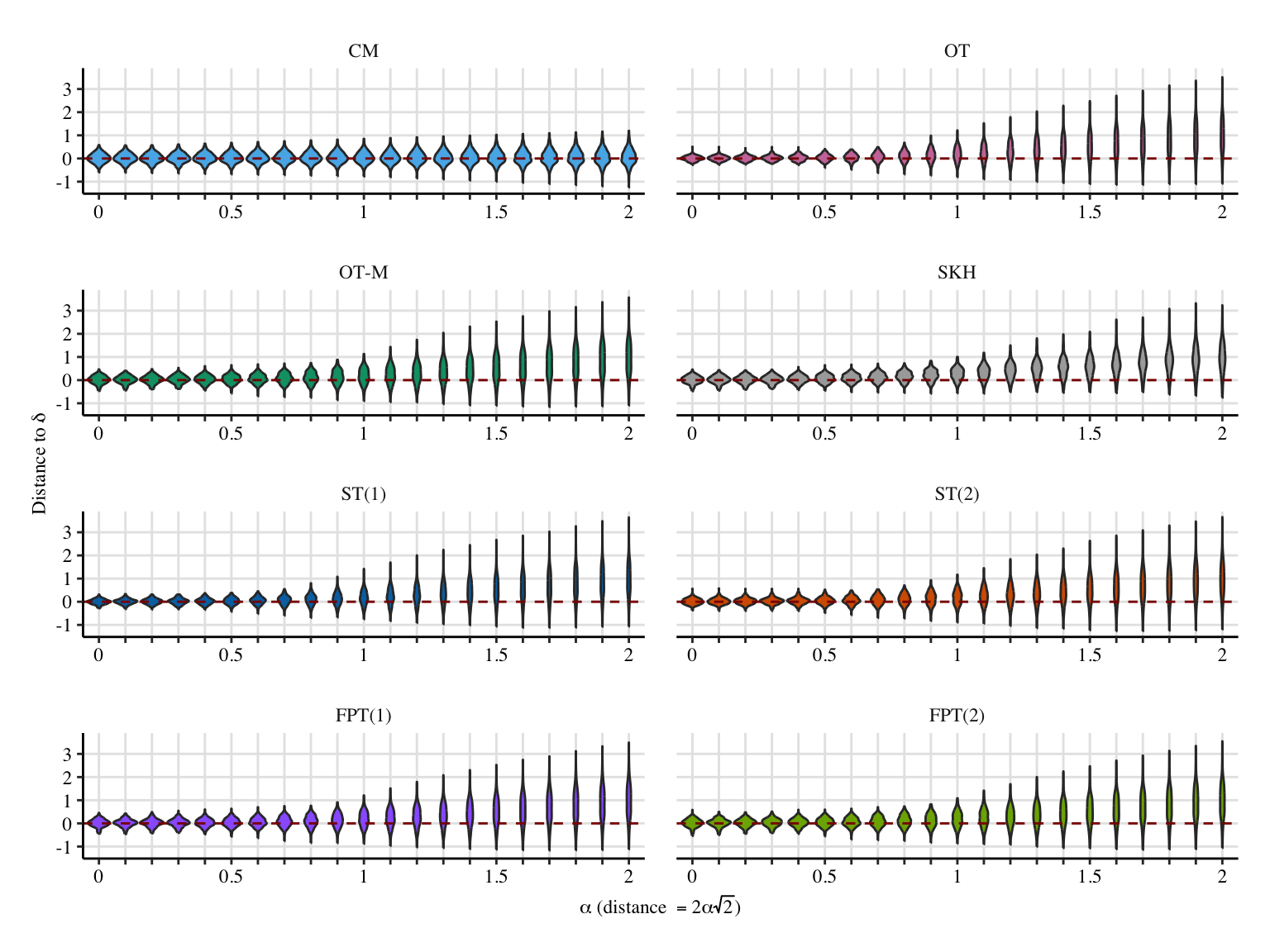
Codes to create the Figure.
export_pdf <- FALSE
x_lab <- "$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"
# y_lab <- "Distance to $\\bar\\zeta(1)$"
y_lab <- "Distance to $\\bar\\zeta$"
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
}
data_plot <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
zeta_1_med, zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
zeta_1_med, zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"zeta_1_med", "zeta_1_ot", "zeta_1_tmatch",
"zeta_1_skh", "zeta_1_sot_12", "zeta_1_sot_21",
"zeta_1_fairadapt_12_lin", "zeta_1_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = 3 - val
)
p <- ggplot(
data = data_plot,
mapping = aes(x = factor(a), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
if (export_pdf == TRUE) {
filename <- "gaussian-mc-alpha-zeta1"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
width = 3.3, height = 3.5,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p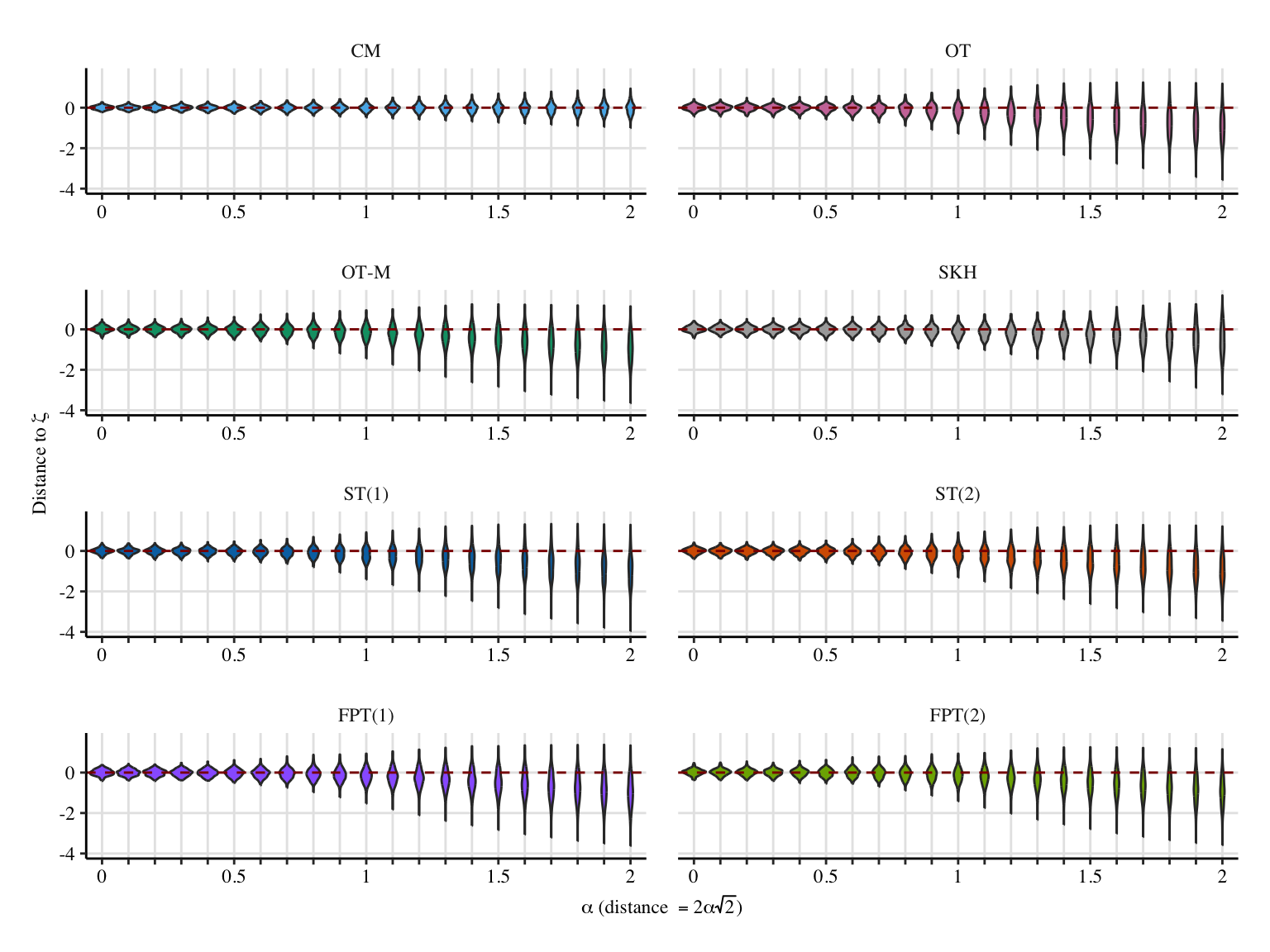
A more condensed figure, which focuses on \(\bar{\delta}(0)\), \(\bar{\zeta}(1)\), and \(\bar{\tau}\).
Codes to create the Figure.
a1 <- 2
a2 <- -1.5
x_lab <- "$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"
y_lab <- "Estimation error"
metrics_lab <- c("$\\bar{\\delta}(0)$", "$\\bar{\\zeta}(1)$", "$\\bar{\\tau}$")
metrics_lab <- c("$\\bar{\\delta}$", "$\\bar{\\zeta}$", "$\\bar{\\tau}$")
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
metrics_lab <- latex2exp::TeX(metrics_lab)
data_plot_tau <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
tot_effect_med,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin,
tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21
) |>
pivot_longer(
cols = c(
tot_effect_med,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin,
tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21
),
names_to = "type", values_to = "tau"
) |>
mutate(
type = factor(
type,
levels = c(
"tot_effect_med",
"tot_effect_fairadapt_12_lin", "tot_effect_fairadapt_21_lin",
"tot_effect_ot", "tot_effect_tmatch",
"tot_effect_skh", "tot_effect_sot_12", "tot_effect_sot_21"
),
labels = c("CM", "FPT(1)", "FPT(2)", "OT", "OT-M", "SKH", "ST(1)", "ST(2)")
),
dist_to_causal_effect = (a1+a2) * (a*mu1 - a*mu0) + 3 - tau
)
data_plot_delta0 <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
delta_0_med,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin,
delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21
) |>
pivot_longer(
cols = c(
delta_0_med,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin,
delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21
),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"delta_0_med",
"delta_0_fairadapt_12_lin", "delta_0_fairadapt_21_lin",
"delta_0_ot", "delta_0_tmatch",
"delta_0_skh", "delta_0_sot_12", "delta_0_sot_21"
),
labels = c("CM", "FPT(1)", "FPT(2)", "OT", "OT-M", "SKH", "ST(1)", "ST(2)")
),
dist_to_theo_val = ((a1+a2)*(a*mu1-a*mu0)) - val
)
data_plot_zeta1 <- res_sim |>
dplyr::select(
n0, n1, seed, a, mu0, mu1,
zeta_1_med,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin,
zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21
) |>
pivot_longer(
cols = c(
zeta_1_med, zeta_1_ot, zeta_1_tmatch,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"zeta_1_med",
"zeta_1_fairadapt_12_lin", "zeta_1_fairadapt_21_lin",
"zeta_1_ot", "zeta_1_tmatch",
"zeta_1_skh", "zeta_1_sot_12", "zeta_1_sot_21"
),
labels = c("CM", "FPT(1)", "FPT(2)", "OT", "OT-M", "SKH", "ST(1)", "ST(2)")
),
dist_to_theo_val = 3 - val
)
data_plot <- data_plot_tau |>
rename(dist_tau = dist_to_causal_effect) |>
left_join(
data_plot_delta0 |>
rename(
delta0 = val,
dist_delta0 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type")
) |>
left_join(
data_plot_zeta1 |>
rename(
zeta1 = val,
dist_zeta1 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type")
) |>
filter(a %in% seq(0, 2, by = .2))
p <- ggplot(
data = data_plot |> select(-tau, -delta0, -zeta1) |>
pivot_longer(
cols = c(dist_tau, dist_delta0, dist_zeta1),
names_to = "metric",
values_to = "dist"
) |>
mutate(
metric = factor(
metric,
levels = c("dist_delta0", "dist_zeta1", "dist_tau"),
labels = metrics_lab
)
),
mapping = aes(x = factor(a), y = dist)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_grid(metric ~ type, scales = "free_x") +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]],
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]]
),
guide = "none",
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
p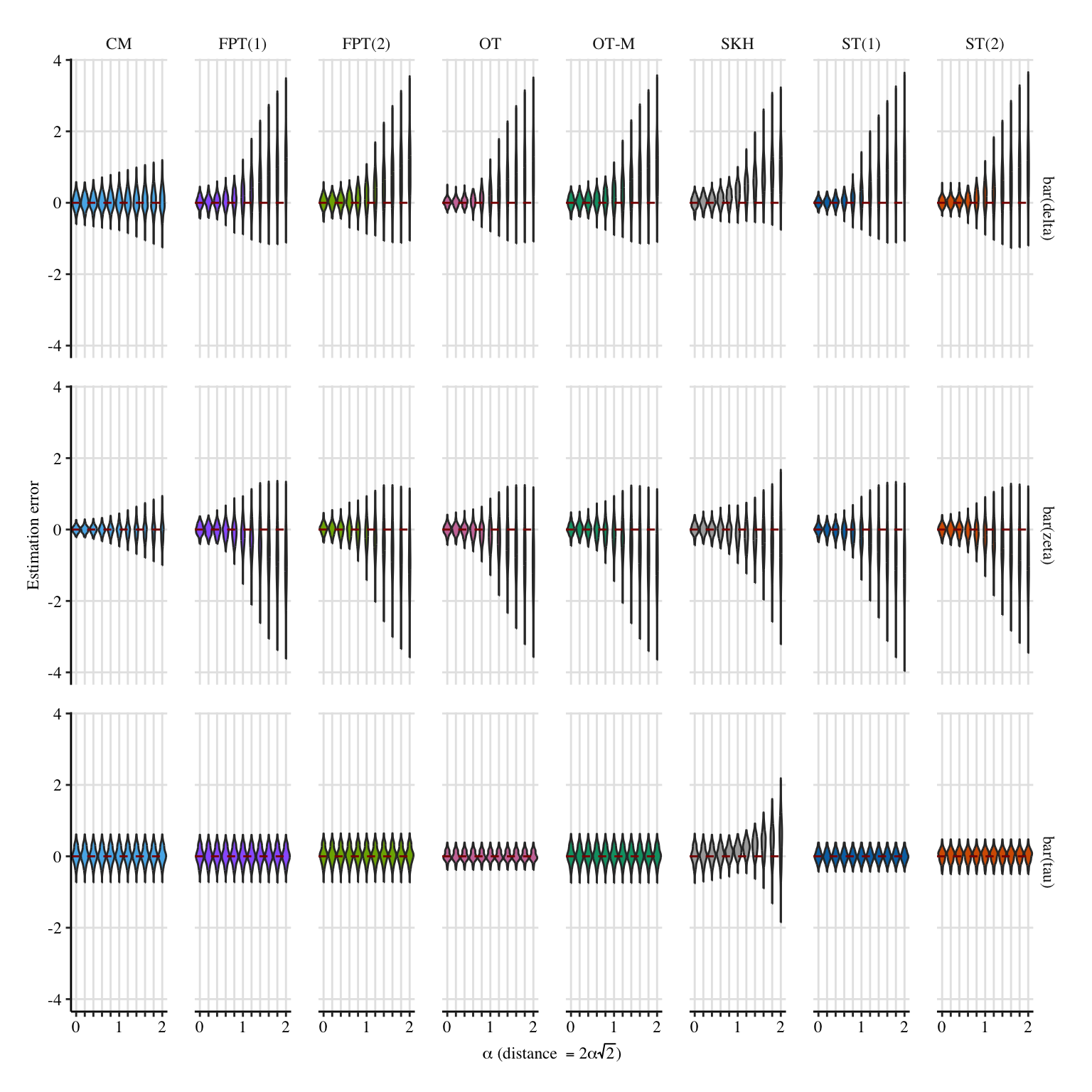
A shorter version for the paper.
Codes to create the Figure.
a1 <- 2
a2 <- -1.5
export_pdf <- TRUE #FALSE
x_lab <- "$\\alpha$ (distance $=2\\alpha\\sqrt{2}$)"
y_lab <- "Estimation error"
metrics_lab <- c("$\\bar{\\delta}(0)$", "$\\bar{\\zeta}(1)$", "$\\bar{\\tau}$")
metrics_lab <- c("$\\bar{\\delta}$", "$\\bar{\\zeta}$", "$\\bar{\\tau}$")
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
metrics_lab <- latex2exp::TeX(metrics_lab)
}
data_plot <- data_plot_tau |>
rename(dist_tau = dist_to_causal_effect) |>
left_join(
data_plot_delta0 |>
rename(
delta0 = val,
dist_delta0 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type")
) |>
left_join(
data_plot_zeta1 |>
rename(
zeta1 = val,
dist_zeta1 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type")
) |>
filter(a %in% seq(0, 2, by = .2)) |>
filter(type %in% c("ST(1)", "ST(2)", "OT", "SKH", "OT"))
p <- ggplot(
data = data_plot |> select(-tau, -delta0, -zeta1) |>
pivot_longer(
cols = c(dist_tau, dist_delta0, dist_zeta1),
names_to = "metric",
values_to = "dist"
) |>
mutate(
metric = factor(
metric,
levels = c("dist_delta0", "dist_zeta1", "dist_tau"),
labels = metrics_lab
)
),
mapping = aes(x = factor(a), y = dist)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_grid(metric ~ type, scales = "free_x") +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]],
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]]
),
guide = "none",
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = ifelse(unique(data_plot$a) %% .5 == 0, unique(data_plot$a), "")
)
scale <- .77
if (export_pdf == TRUE) {
filename <- "gaussian-mc-alpha"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
height = scale*3.3, width = scale*6,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p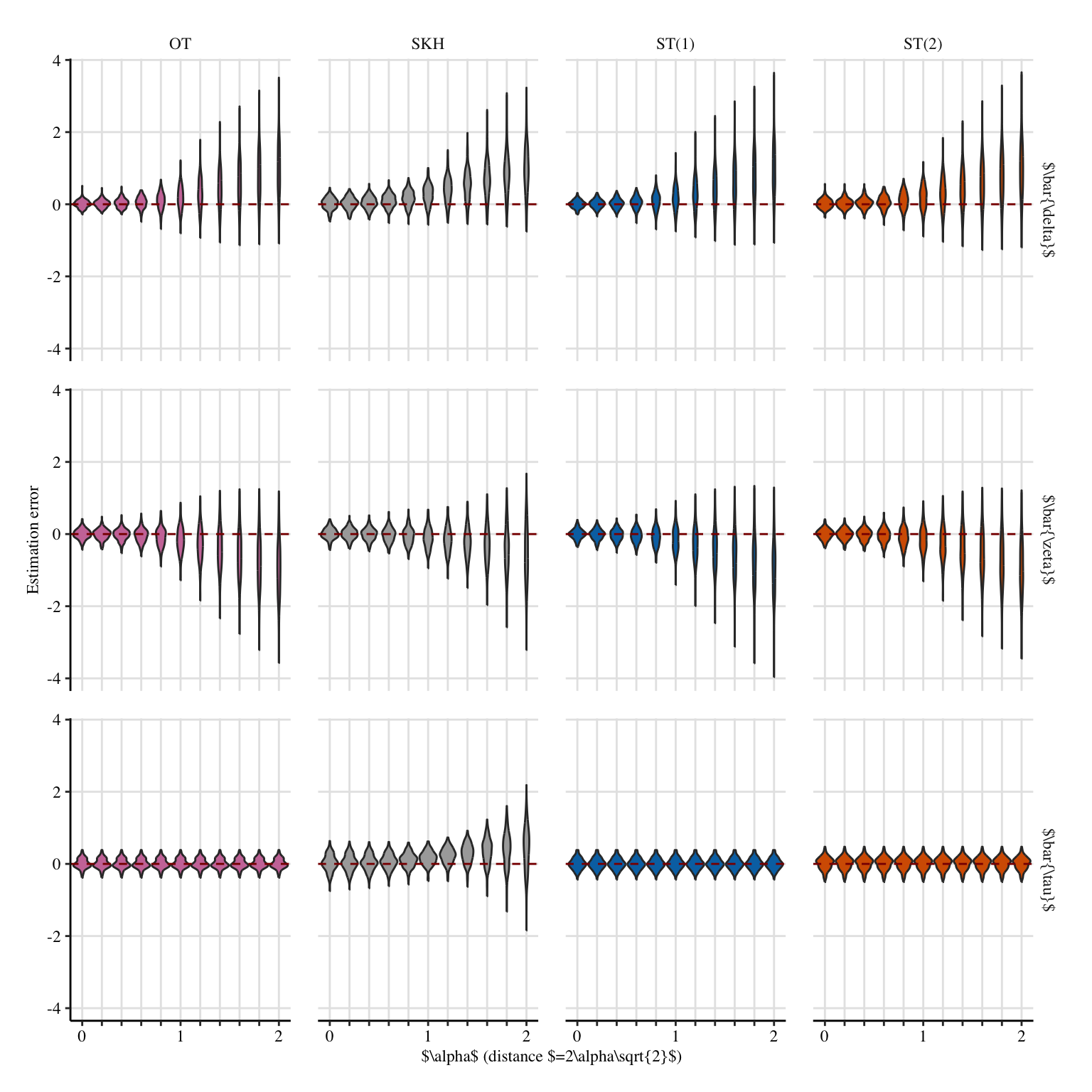
6.3 Varying the Proportion of 0s and 1s
In this section, we let the proportion of untreated/treated units vary from 5% of 0s to 95% of them. Note that we go back to the distance between the mean equal to 1, as in Chapter 5.
A visual representation of the samples that can be drawn from the DGP with different proportions 0s is shown in Figure 6.2
Codes to create the Figure.
set.seed(123)
alpha <- 1
mu0 <- -1
mu1 <- +1
r0 <- +.7
r1 <- -.5
# Generate data with alpha=0, alpha=1, alpha=2
data_alpha_10 <- gen_data(
a = alpha, mu0 = mu0, mu1 = mu1, r0 = r0, r1 = r1,
n0 = 50, n1 = 450
)
data_alpha_50 <- gen_data(
a = alpha, mu0 = mu0, mu1 = mu1, r0 = r0, r1 = r1,
n0 = 25, n1 = 475
)
data_alpha_90 <- gen_data(
a = alpha, mu0 = mu0, mu1 = mu1, r0 = r0, r1 = r1,
n0 = 450, n1 = 50
)
par(mar = c(2.1, 2.1, 2.1, 0.1), mfrow = c(1, 3))
x_lim <- c(-4, 4)
y_lim <- c(-4, 4)
cex_pts <- .4
# With 10%----
a <- 0
x_lab <- "Prop. 0s: $10\\%$"
if (export_tikz == FALSE) x_lab <- latex2exp::TeX(x_lab)
plot(
data_alpha_10$X1[data_alpha_10$A == 0],
data_alpha_10$X2[data_alpha_10$A == 0],
pch = 16,
cex = cex_pts,
col = adjustcolor(colGpe0, alpha = .3),
xlim = x_lim, ylim = y_lim,
xlab = "", ylab = "",
main = x_lab,
family = font_family,
axes = FALSE
)
axis(1, at = -3:3)
axis(2, at = -3:3)
title(xlab = "X1", ylab="X2", line=2, cex.lab=1.2, family = font_family)
points(
data_alpha_10$X1[data_alpha_10$A == 1],
data_alpha_10$X2[data_alpha_10$A == 1],
col = adjustcolor(colGpe1, alpha = .3), pch = 16, cex = cex_pts
)
# True mean and covariance (scaled by 'a')
Mu0 <- rep(alpha * mu0, 2)
Mu1 <- rep(alpha * mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Add ellipses
draw_ellipse(Mu0, Sig0, col = colGpe0, lty = 2)
draw_ellipse(Mu1, Sig1, col = colGpe1, lty = 2)
# With 50%----
a <- 1
x_lab <- "Prop. 0s: $50\\%$"
if (export_tikz == FALSE) x_lab <- latex2exp::TeX(x_lab)
plot(
data_alpha_50$X1[data_alpha_50$A == 0],
data_alpha_50$X2[data_alpha_50$A == 0],
pch = 16, cex = cex_pts,
col = adjustcolor(colGpe0, alpha = .3),
xlim = x_lim, ylim = y_lim,
xlab = "", ylab = "",
main = x_lab,
family = font_family,
axes = FALSE
)
axis(1, at = -3:3)
axis(2, at = -3:3)
title(xlab = "X1", ylab="X2", line=2, cex.lab=1.2, family = font_family)
points(
data_alpha_50$X1[data_alpha_50$A == 1],
data_alpha_50$X2[data_alpha_50$A == 1],
col = adjustcolor(colGpe1, alpha = .3), pch = 16, cex = cex_pts
)
# True mean and covariance (scaled by 'a')
Mu0 <- rep(alpha * mu0, 2)
Mu1 <- rep(alpha * mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Add ellipses
draw_ellipse(Mu0, Sig0, col = colGpe0, lty = 2)
draw_ellipse(Mu1, Sig1, col = colGpe1, lty = 2)
# With 90%----
a <- 2
x_lab <- "Prop. 0s: $90\\%$"
if (export_tikz == FALSE) x_lab <- latex2exp::TeX(x_lab)
plot(
data_alpha_90$X1[data_alpha_90$A == 0],
data_alpha_90$X2[data_alpha_90$A == 0],
pch = 16, cex = cex_pts,
col = adjustcolor(colGpe0, alpha = .3),
xlim = x_lim, ylim = y_lim,
xlab = "", ylab = "",
main = x_lab,
family = font_family,
axes = FALSE
)
axis(1, at = -3:3)
axis(2, at = -3:3)
title(xlab = "X1", ylab="X2", line=2, cex.lab=1.2, family = font_family)
points(
data_alpha_90$X1[data_alpha_90$A == 1],
data_alpha_90$X2[data_alpha_90$A == 1],
col = adjustcolor(colGpe1, alpha = .3), pch = 16, cex = cex_pts
)
# True mean and covariance (scaled by 'a')
Mu0 <- rep(alpha * mu0, 2)
Mu1 <- rep(alpha * mu1, 2)
Sig0 <- matrix(c(1, r0, r0, 1), 2, 2)
Sig1 <- matrix(c(1, r1, r1, 1), 2, 2)
# Add ellipses
draw_ellipse(Mu0, Sig0, col = colGpe0, lty = 2)
draw_ellipse(Mu1, Sig1, col = colGpe1, lty = 2)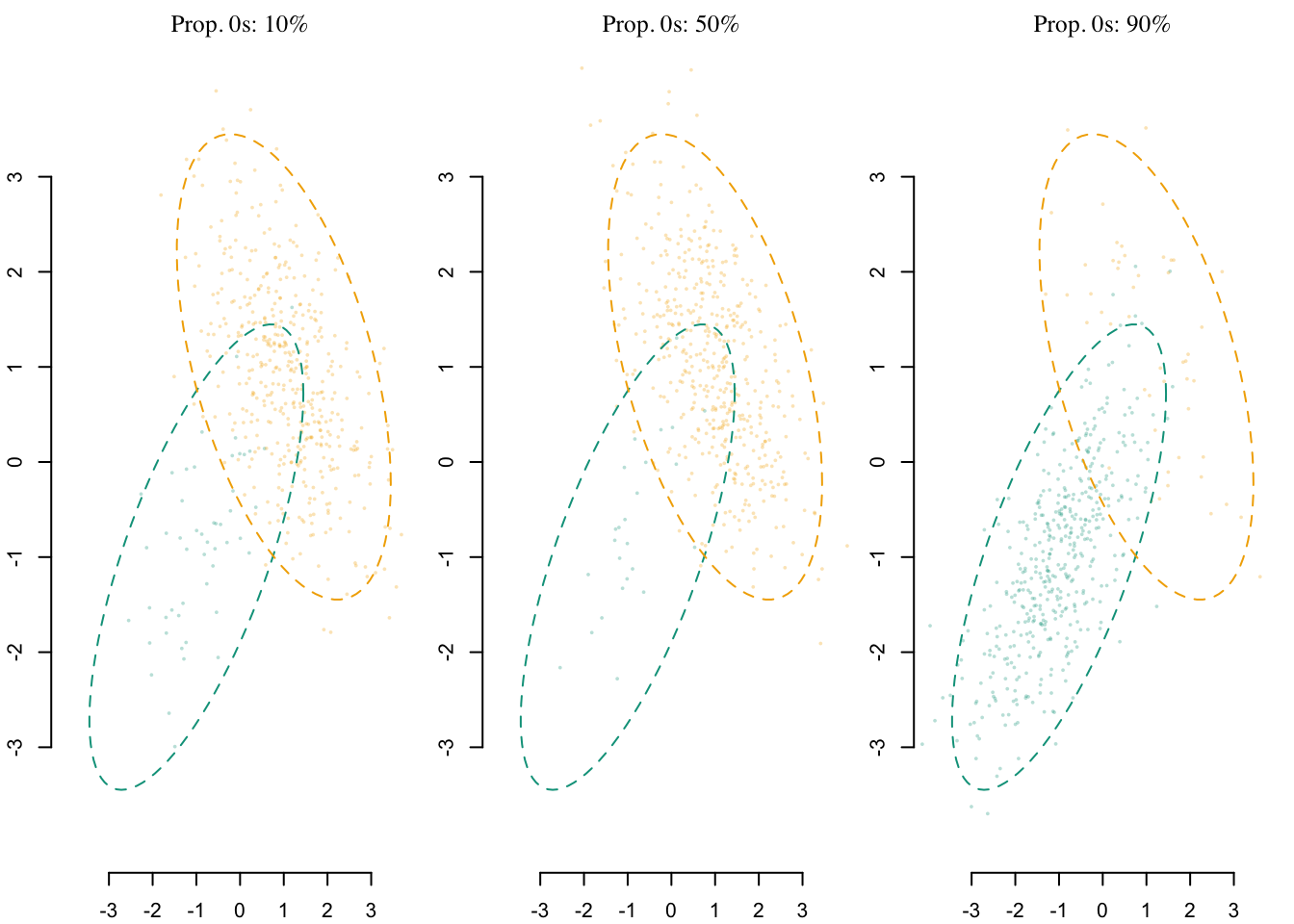
We define a grid with the different simulations.
grid_params <- expand_grid(
prop_0 = seq(5, 95, by = 5)/100,
mu0 = -1,
mu1 = +1,
r0 = +.7,
r1 = -.5,
a = 1,
seed = seq_len(n_repl)
)The simulations can be run in parallel, as follows.
The codes to run the simulations
# This chunk takes about XX minutes to run.
# We do not evaluate when compiling the document.
# Instead, we load previously obtained results.
library(pbapply)
library(parallel)
ncl <- detectCores()-1
(cl <- makeCluster(ncl))
clusterEvalQ(cl, {
library(tidyverse)
library(mnormt)
library(expm)
library(randomForest)
library(fairadapt)
}) |>
invisible()
clusterExport(
cl = cl, c(
"gen_data", "compute_ot_map", "apply_ot_transport",
"transport_regul", "transport_many_to_one",
"sequential_transport_12", "sequential_transport_21",
"causal_effects_cf", "sim_f", "grid_params",
"fairadapt_12", "fairadapt_21"
)
)
res_sim_prop <- pbapply::pblapply(
1:nrow(grid_params), function(i) {
grid_params_current <- grid_params[i, ]
n <- 500
prop_0 <- grid_params_current$prop_0
n0 <- prop_0 * n
n1 <- n - n0
mu0 <- grid_params_current$mu0
mu1 <- grid_params_current$mu1
r0 <- grid_params_current$r0
r1 <- grid_params_current$r1
a <- grid_params_current$a
seed <- grid_params_current$seed
sim_f(
n0 = n0, n1 = n1,
mu0 = mu0, mu1 = mu1,
r0 = r0, r1 = r1,
a = a, seed = seed
) |>
mutate(prop_0 = prop_0)
}, cl = cl)
stopCluster(cl)
res_sim_prop <- list_rbind(res_sim_prop)
save(
res_sim_prop,
file = "../output/res_sim_prop-gaussian-mc.rda"
)We load previously obtained results:
load("../output/res_sim_prop-gaussian-mc.rda")Codes to create the Figure.
a1 <- 2
a2 <- -1.5
export_pdf <- TRUE #FALSE
x_lab <- "Proportion of untreated"
y_lab <- "Distance to $\\bar\\tau$"
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
}
data_plot <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
tot_effect_med, tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
tot_effect_med, tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin
),
names_to = "type", values_to = "tau"
) |>
mutate(
type = factor(
type,
levels = c(
"tot_effect_med", "tot_effect_ot", "tot_effect_tmatch",
"tot_effect_skh", "tot_effect_sot_12", "tot_effect_sot_21",
"tot_effect_fairadapt_12_lin", "tot_effect_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_causal_effect = (a1+a2) * (a*mu1 - a*mu0) + 3 - tau
)
p <- ggplot(
data = data_plot,
mapping = aes(x = factor(prop_0), y = dist_to_causal_effect)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none",
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = ifelse(
(unique(data_plot$prop_0)*100) %% 20 == 0,
yes = unique(data_plot$prop_0), no = "")
)
if (export_pdf == TRUE) {
filename <- "gaussian-mc-prop-tau"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
width = 3.3, height = 3.5,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p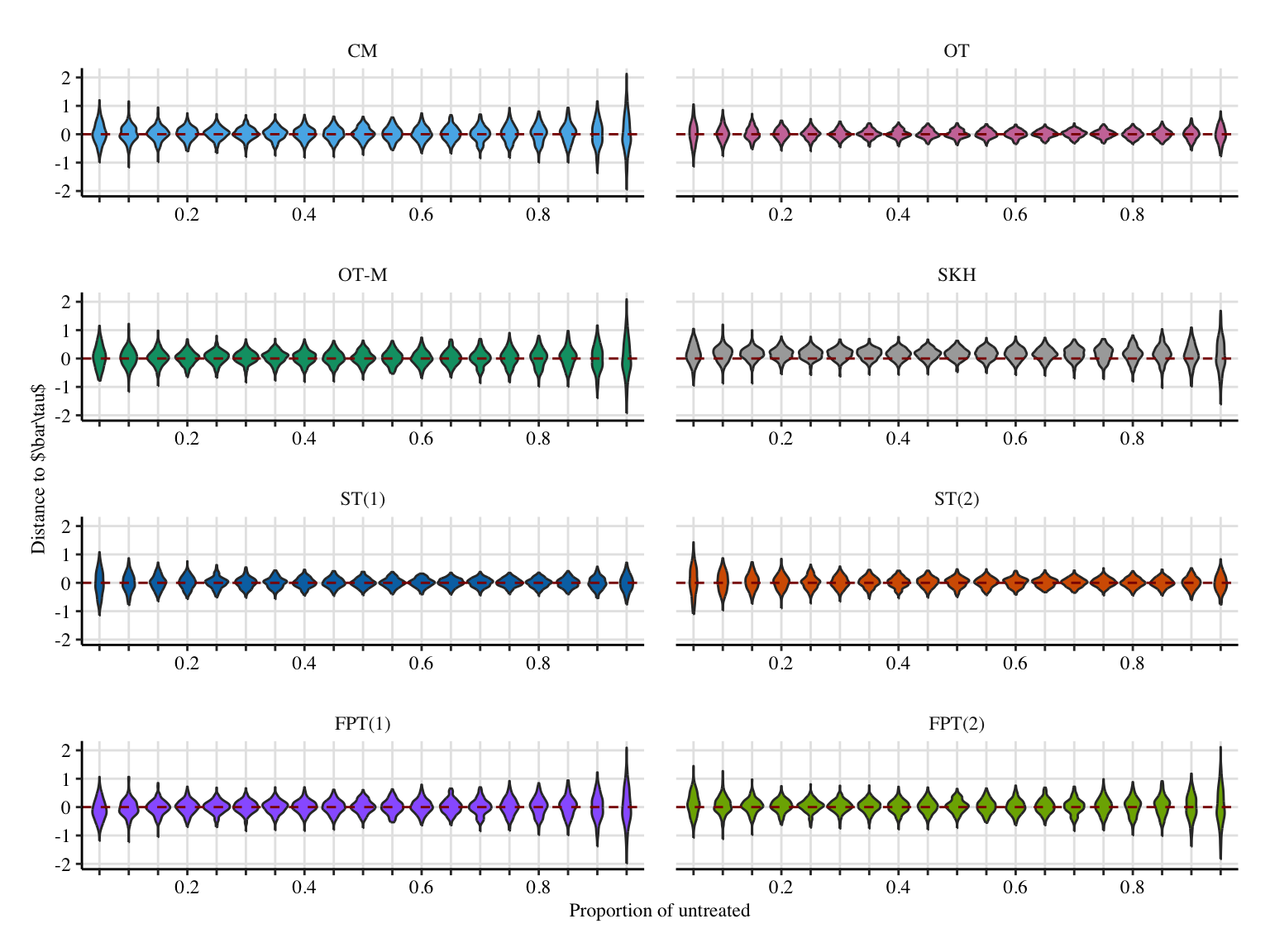
Codes to create the Figure.
a1 <- 2
a2 <- -1.5
export_pdf <- FALSE
x_lab <- "Proportion of untreated"
# y_lab <- "Distance to $\\bar\\delta(0)$"
y_lab <- "Distance to $\\bar\\delta$"
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
}
data_plot <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
delta_0_med, delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
delta_0_med, delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin
),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"delta_0_med", "delta_0_ot", "delta_0_tmatch",
"delta_0_skh", "delta_0_sot_12", "delta_0_sot_21",
"delta_0_fairadapt_12_lin", "delta_0_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = ((a1+a2)*(mu1-mu0)) - val
)
p <- ggplot(
data = data_plot,
mapping = aes(x = factor(prop_0), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab
) +
scale_x_discrete(
labels = ifelse(
(unique(data_plot$prop_0)*100) %% 20 == 0,
yes = unique(data_plot$prop_0), no = "")
)
if (export_pdf == TRUE) {
filename <- "gaussian-mc-prop-delta0"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
width = 3.3, height = 3.5,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p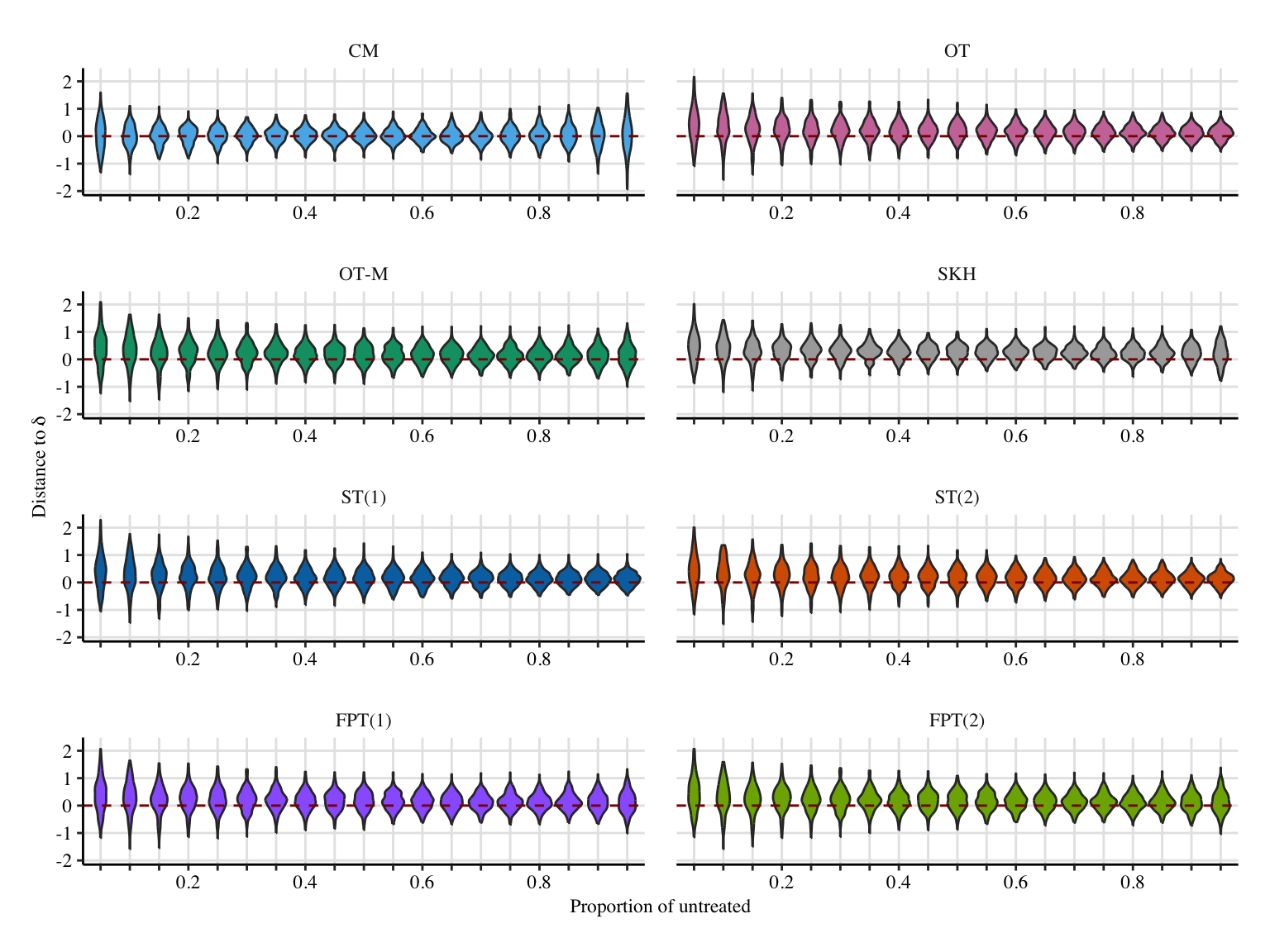
Codes to create the Figure.
data_plot <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
delta_1_med, delta_1_ot, delta_1_tmatch,
delta_1_skh, delta_1_sot_12, delta_1_sot_21,
delta_1_fairadapt_12_lin, delta_1_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
delta_1_med, delta_1_ot, delta_1_tmatch,
delta_1_skh, delta_1_sot_12, delta_1_sot_21,
delta_1_fairadapt_12_lin, delta_1_fairadapt_21_lin),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"delta_1_med", "delta_1_ot", "delta_1_tmatch",
"delta_1_skh", "delta_1_sot_12", "delta_1_sot_21",
"delta_1_fairadapt_12_lin", "delta_1_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = ((a1+a2)*(mu1-mu0)) - val
)
ggplot(
data = data_plot,
mapping = aes(x = factor(prop_0), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
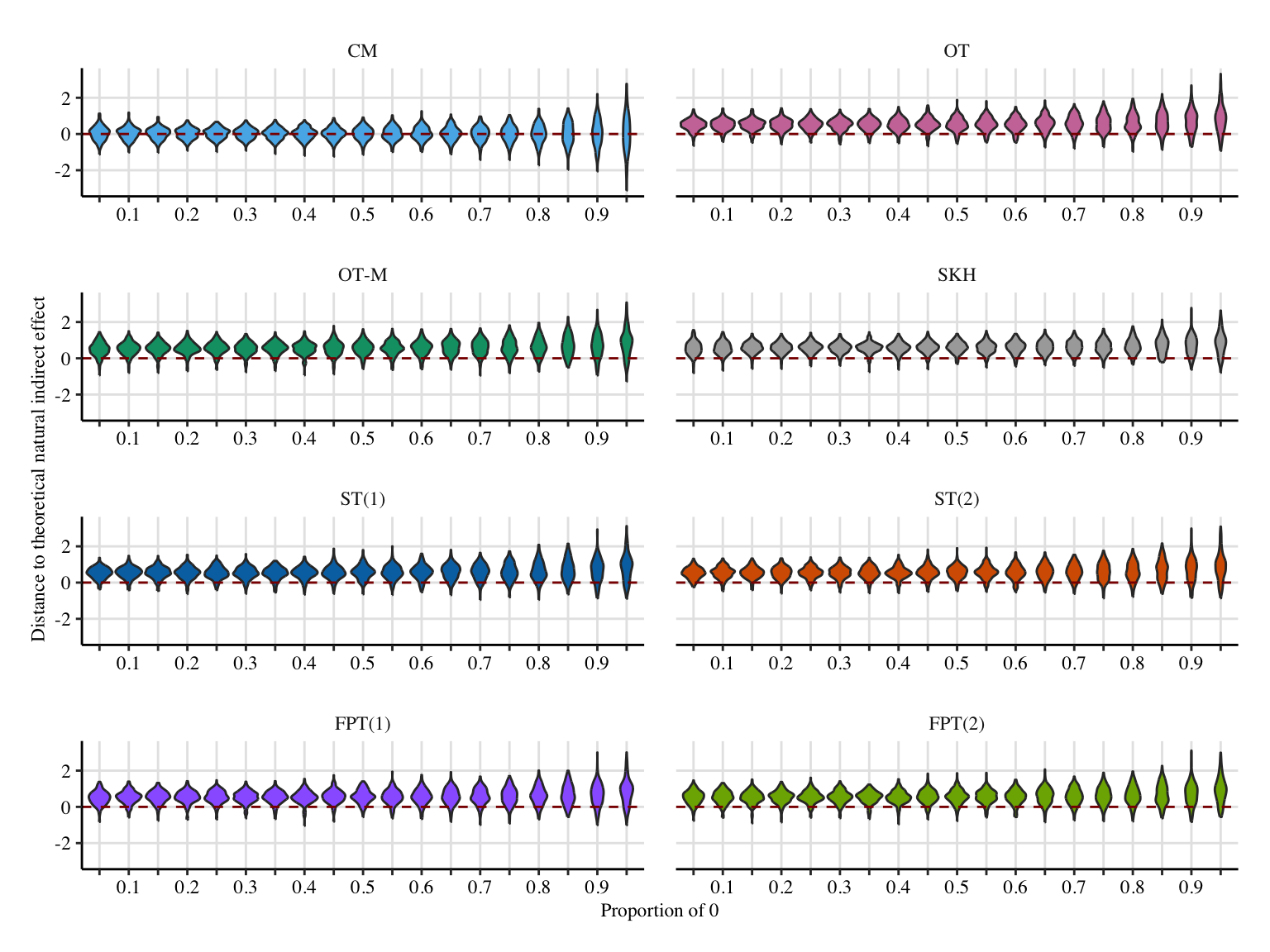
labs(
x = "Proportion of 0",
y = "Distance to theoretical natural indirect effect",
) +
scale_x_discrete(
labels = ifelse(
(unique(data_plot$prop_0)*100) %% 10 == 0,
yes = unique(data_plot$prop_0), no = "")
)
Codes to create the Figure.
data_plot <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
zeta_0_med, zeta_0_ot, zeta_0_tmatch,
zeta_0_skh, zeta_0_sot_12, zeta_0_sot_21,
zeta_0_fairadapt_12_lin, zeta_0_fairadapt_21_lin
) |>
pivot_longer(
cols = c(zeta_0_med, zeta_0_ot, zeta_0_tmatch,
zeta_0_skh, zeta_0_sot_12, zeta_0_sot_21,
zeta_0_fairadapt_12_lin, zeta_0_fairadapt_21_lin),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"zeta_0_med", "zeta_0_ot", "zeta_0_tmatch",
"zeta_0_skh", "zeta_0_sot_12", "zeta_0_sot_21",
"zeta_0_fairadapt_12_lin", "zeta_0_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = 3 - val
)
ggplot(
data = data_plot,
mapping = aes(x = factor(prop_0), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
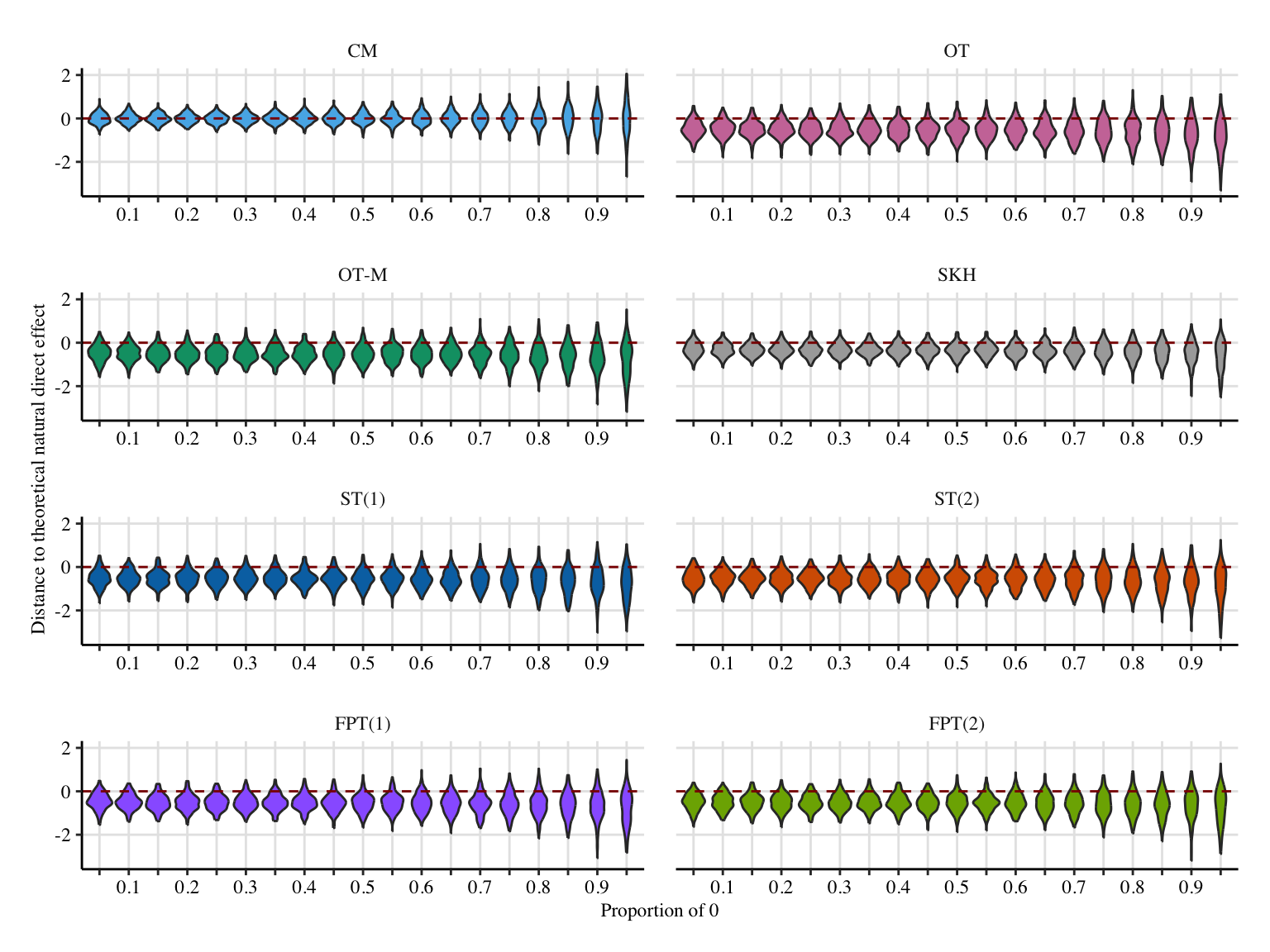
labs(
x = "Proportion of 0",
y = "Distance to theoretical natural direct effect",
) +
scale_x_discrete(
labels = ifelse(
(unique(data_plot$prop_0)*100) %% 10 == 0,
yes = unique(data_plot$prop_0), no = "")
)
Codes to create the Figure.
a1 <- 2
a2 <- -1.5
export_pdf <- FALSE
x_lab <- "Proportion of untreated"
# y_lab <- "Distance to $\\bar\\zeta(1)$"
y_lab <- "Distance to $\\bar\\zeta$"
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
}
data_plot <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
zeta_1_med, zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin
) |>
pivot_longer(
cols = c(
zeta_1_med, zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"zeta_1_med", "zeta_1_ot", "zeta_1_tmatch",
"zeta_1_skh", "zeta_1_sot_12", "zeta_1_sot_21",
"zeta_1_fairadapt_12_lin", "zeta_1_fairadapt_21_lin"
),
labels = c("CM", "OT", "OT-M", "SKH", "ST(1)", "ST(2)", "FPT(1)", "FPT(2)")
),
dist_to_theo_val = 3 - val
)
p <- ggplot(
data = data_plot,
mapping = aes(x = factor(prop_0), y = dist_to_theo_val)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_wrap(~type, scales = "free_x", ncol = 2) +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]],
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]]
),
guide = "none"
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab
) +
scale_x_discrete(
labels = ifelse(
(unique(data_plot$prop_0)*100) %% 20 == 0,
yes = unique(data_plot$prop_0), no = "")
)
if (export_pdf == TRUE) {
filename <- "gaussian-mc-prop-zeta1"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")),
filename = filename, path = "figs/",
width = 3.3, height = 3.5,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p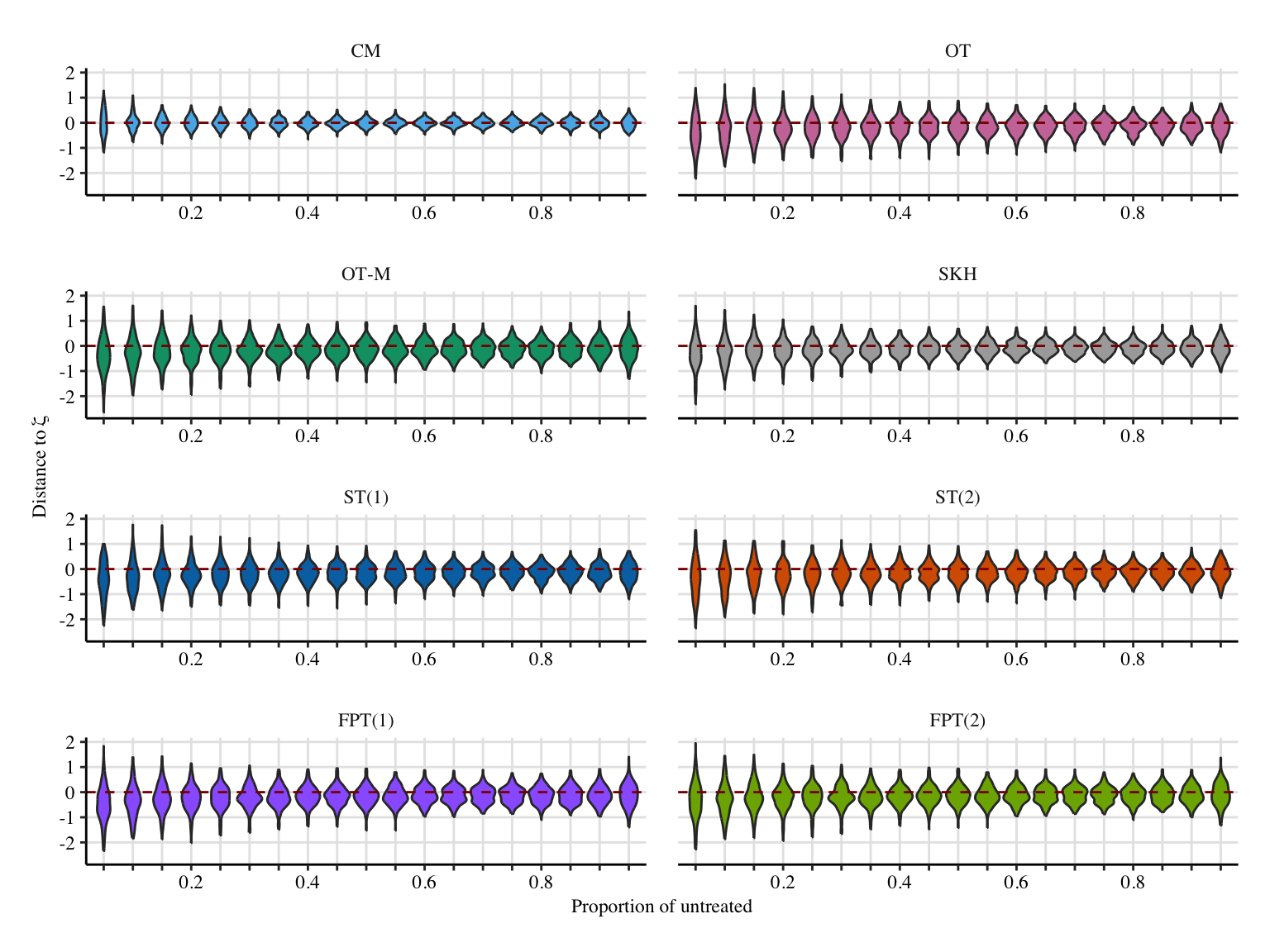
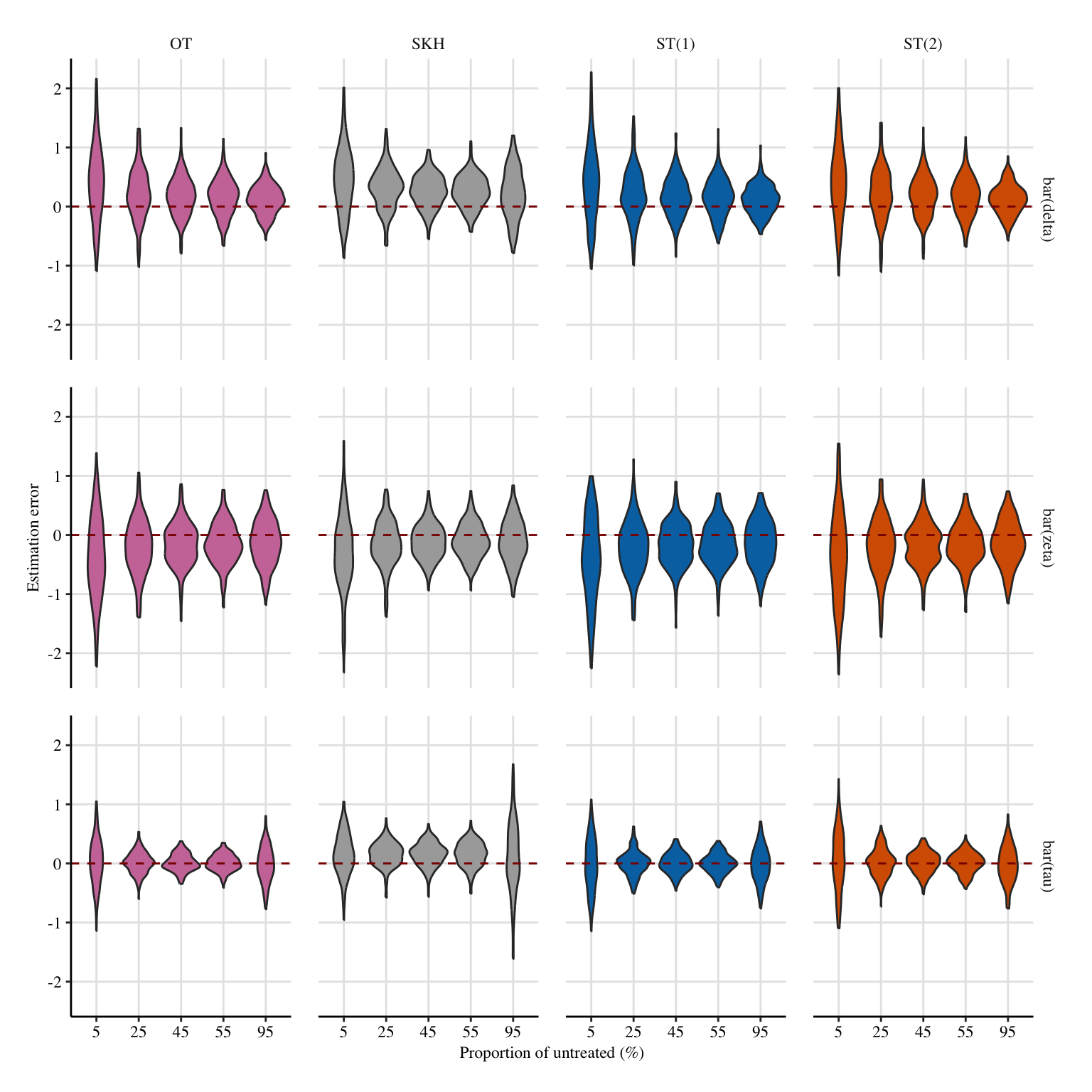
A more condensed figure, which focuses on \(\bar{\delta}(0)\), \(\bar{\zeta}(1)\), and \(\bar{\tau}\).
Codes to create the Figure.
a1 <- 2
a2 <- -1.5
x_lab <- "Proportion of untreated ($\\%$)"
y_lab <- "Estimation error"
# metrics_lab <- c("$\\bar{\\delta}(0)$", "$\\bar{\\zeta}(1)$", "$\\bar{\\tau}$")
metrics_lab <- c("$\\bar{\\delta}$", "$\\bar{\\zeta}$", "$\\bar{\\tau}$")
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
metrics_lab <- latex2exp::TeX(metrics_lab)
data_plot_tau <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
tot_effect_med,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin,
tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21
) |>
pivot_longer(
cols = c(
tot_effect_med,
tot_effect_fairadapt_12_lin, tot_effect_fairadapt_21_lin,
tot_effect_ot, tot_effect_tmatch,
tot_effect_skh, tot_effect_sot_12, tot_effect_sot_21
),
names_to = "type", values_to = "tau"
) |>
mutate(
type = factor(
type,
levels = c(
"tot_effect_med",
"tot_effect_fairadapt_12_lin", "tot_effect_fairadapt_21_lin",
"tot_effect_ot", "tot_effect_tmatch",
"tot_effect_skh", "tot_effect_sot_12", "tot_effect_sot_21"
),
labels = c("CM", "FPT(1)", "FPT(2)", "OT", "OT-M", "SKH", "ST(1)", "ST(2)")
),
dist_to_causal_effect = (a1+a2) * (a*mu1 - a*mu0) + 3 - tau
)
data_plot_delta0 <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
delta_0_med,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin,
delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21
) |>
pivot_longer(
cols = c(
delta_0_med,
delta_0_fairadapt_12_lin, delta_0_fairadapt_21_lin,
delta_0_ot, delta_0_tmatch,
delta_0_skh, delta_0_sot_12, delta_0_sot_21
),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"delta_0_med",
"delta_0_fairadapt_12_lin", "delta_0_fairadapt_21_lin",
"delta_0_ot", "delta_0_tmatch",
"delta_0_skh", "delta_0_sot_12", "delta_0_sot_21"
),
labels = c("CM", "FPT(1)", "FPT(2)", "OT", "OT-M", "SKH", "ST(1)", "ST(2)")
),
dist_to_theo_val = ((a1+a2)*(mu1-mu0)) - val
)
data_plot_zeta1 <- res_sim_prop |>
dplyr::select(
n0, n1, seed, a, mu0, mu1, prop_0,
zeta_1_med,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin,
zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21
) |>
pivot_longer(
cols = c(
zeta_1_med,
zeta_1_fairadapt_12_lin, zeta_1_fairadapt_21_lin,
zeta_1_ot, zeta_1_tmatch,
zeta_1_skh, zeta_1_sot_12, zeta_1_sot_21),
names_to = "type", values_to = "val"
) |>
mutate(
type = factor(
type,
levels = c(
"zeta_1_med",
"zeta_1_fairadapt_12_lin", "zeta_1_fairadapt_21_lin",
"zeta_1_ot", "zeta_1_tmatch",
"zeta_1_skh", "zeta_1_sot_12", "zeta_1_sot_21"
),
labels = c("CM", "FPT(1)", "FPT(2)", "OT", "OT-M", "SKH", "ST(1)", "ST(2)")
),
dist_to_theo_val = 3 - val
)
data_plot <- data_plot_tau |>
rename(dist_tau = dist_to_causal_effect) |>
left_join(
data_plot_delta0 |>
rename(
delta0 = val,
dist_delta0 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type", "prop_0")
) |>
left_join(
data_plot_zeta1 |>
rename(
zeta1 = val,
dist_zeta1 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type", "prop_0")
)
data_plot <- data_plot |> filter(prop_0 %in% seq(.05, .95, by =.1))
p <- ggplot(
data = data_plot |> select(-tau, -delta0, -zeta1) |>
pivot_longer(
cols = c(dist_tau, dist_delta0, dist_zeta1),
names_to = "metric",
values_to = "dist"
) |>
mutate(
metric = factor(
metric,
levels = c("dist_delta0", "dist_zeta1", "dist_tau"),
labels = metrics_lab
)
),
mapping = aes(x = factor(prop_0), y = dist)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_grid(metric ~ type, scales = "free_x") +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]],
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]]
),
guide = "none",
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = 100*unique(data_plot$prop_0)
)
p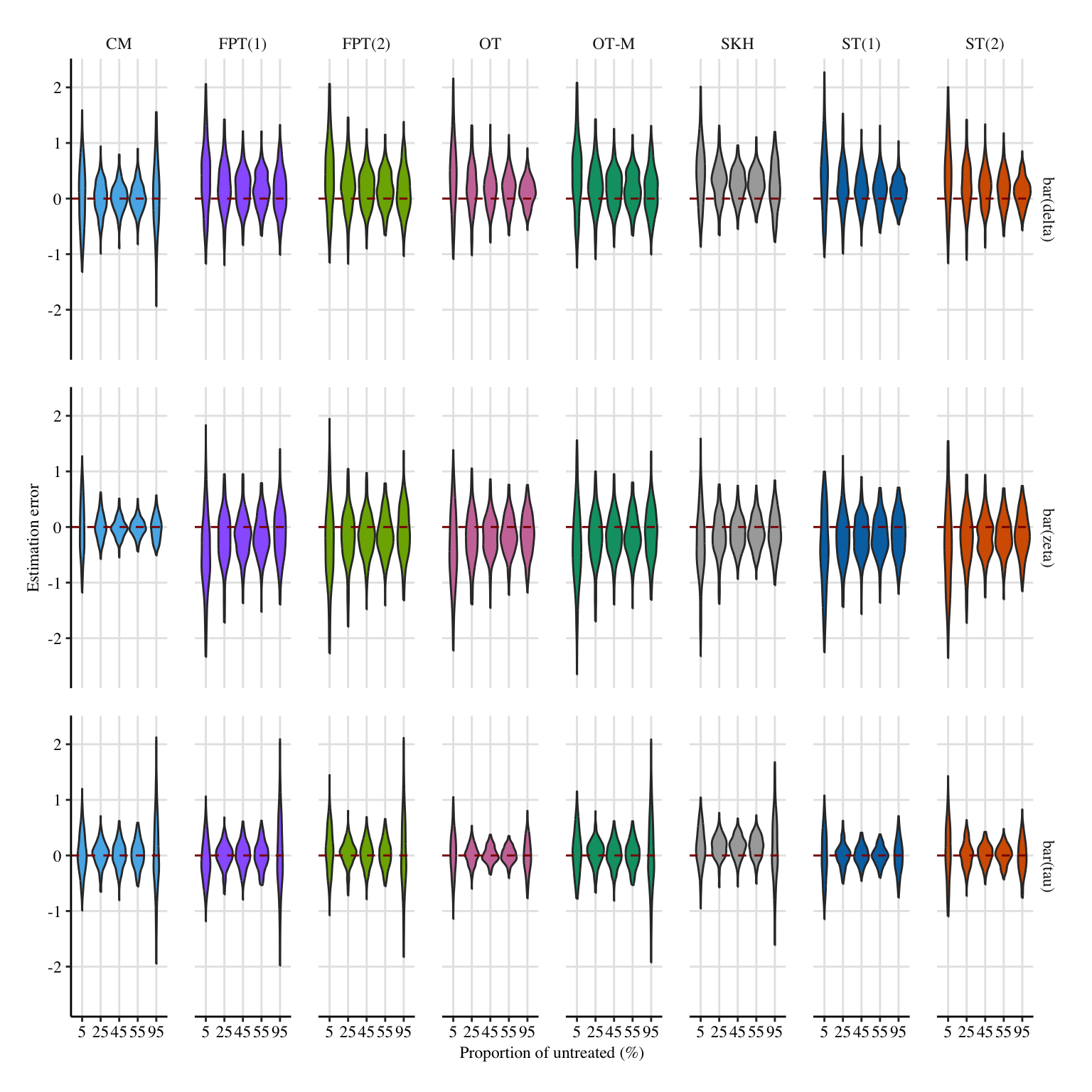
A shorter version of the graph for the paper.
Codes to create the Figure.
a1 <- 2
a2 <- -1.5
export_pdf <- FALSE
x_lab <- "Proportion of untreated ($\\%$)"
y_lab <- "Estimation error"
metrics_lab <- c("$\\bar{\\delta}$", "$\\bar{\\zeta}$", "$\\bar{\\tau}$")
if (export_pdf == FALSE) {
x_lab <- latex2exp::TeX(x_lab)
y_lab <- latex2exp::TeX(y_lab)
metrics_lab <- latex2exp::TeX(metrics_lab)
}
data_plot <- data_plot_tau |>
rename(dist_tau = dist_to_causal_effect) |>
left_join(
data_plot_delta0 |>
rename(
delta0 = val,
dist_delta0 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type", "prop_0")
) |>
left_join(
data_plot_zeta1 |>
rename(
zeta1 = val,
dist_zeta1 = dist_to_theo_val
),
by = c("n0", "n1", "seed", "a", "mu0", "mu1", "type", "prop_0")
)
data_plot <- data_plot |> filter(prop_0 %in% seq(.05, .95, by =.1)) |>
filter(type %in% c("ST(1)", "ST(2)", "OT", "SKH", "OT"))
p <- ggplot(
data = data_plot |> select(-tau, -delta0, -zeta1) |>
pivot_longer(
cols = c(dist_tau, dist_delta0, dist_zeta1),
names_to = "metric",
values_to = "dist"
) |>
mutate(
metric = factor(
metric,
levels = c("dist_delta0", "dist_zeta1", "dist_tau"),
labels = metrics_lab
)
),
mapping = aes(x = factor(prop_0), y = dist)
) +
geom_violin(
mapping = aes(fill = type)
) +
geom_hline(yintercept = 0, colour = "darkred", linetype = "dashed") +
theme_paper() +
facet_grid(metric ~ type, scales = "free_x") +
scale_fill_manual(
NULL,
values = c(
"CM" = "#56B4E9",
"FPT(1)" = colour_methods[["fairadapt_1"]],
"FPT(2)" = colour_methods[["fairadapt_2"]],
"OT" = colour_methods[["OT"]],
"OT-M" = colour_methods[["OT-M"]],
"SKH" = colour_methods[["skh"]],
"ST(1)" = colour_methods[["seq_1"]],
"ST(2)" = colour_methods[["seq_2"]]
),
guide = "none",
) +
theme(legend.position = "bottom") +
labs(
x = x_lab,
y = y_lab,
) +
scale_x_discrete(
labels = 100*unique(data_plot$prop_0)
)
scale <- .77
if (export_pdf == TRUE) {
filename <- "gaussian-mc-prop"
ggplot2_to_pdf(
plot = p + theme(panel.spacing = unit(0.4, "lines")) +
theme(axis.text.x = element_text(size = rel(1))),
filename = filename, path = "figs/",
height = scale*3.6, width = scale*6,
crop = TRUE
)
system(paste0("pdfcrop figs/", filename, ".pdf figs/", filename, ".pdf"))
}
p